Fungi in the Wound Microbiome
- PMID: 29984114
- PMCID: PMC6032664
- DOI: 10.1089/wound.2017.0756
Fungi in the Wound Microbiome
Abstract
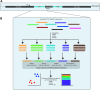
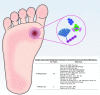
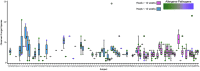
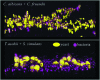
Significance: Culture-independent methods have revealed the diverse and dynamic bacterial communities that colonize chronic wounds. Only recently have studies begun to examine fungal colonization and interactions with the bacterial component of the microbiome, their relationship with the host, and influence on wound outcomes. Recent Advances: Studies using culture-independent sequencing methods reveal that fungi often go undetected in wounds. Candida spp. and Cladosporidium spp. are the most commonly identified fungi in wounds. The wound environment may promote multispecies biofilm formation between bacteria and fungi in wounds, with implications for pathogenicity, treatment, and outcomes. Critical Issues: Identifying microorganisms that are problematic for healing will require a comprehensive understanding of all members of the polymicrobial wound community, including fungi and bacteria. Improved reference databases and bioinformatics tools for studying fungal communities will stimulate further research into the fungal microbiome. Future Directions: Continued study of polymicrobial wound communities using culture-independent methods will further our understanding of the relationships between microbial bioburden, the host response, and impact on healing, complications, and patient outcomes. Future studies should encompass all types of microbiota, including fungi, and focus on potential multi-kingdom interactions that contribute to pathogenicity, biofilm formation, and poor outcomes.
Keywords: biofilm; chronic wounds; fungi; microbiome.
Figures

Similar articles
-
Redefining the Chronic-Wound Microbiome: Fungal Communities Are Prevalent, Dynamic, and Associated with Delayed Healing.mBio. 2016 Sep 6;7(5):e01058-16. doi: 10.1128/mBio.01058-16. mBio. 2016. PMID: 27601572 Free PMC article.
-
Cutaneous wound biofilm and the potential for electrical stimulation in management of the microbiome.Future Microbiol. 2017 Mar;12:337-357. doi: 10.2217/fmb-2016-0204. Epub 2017 Mar 3. Future Microbiol. 2017. PMID: 28287302 Review.
-
The role of the microbiome in nonhealing diabetic wounds.Ann N Y Acad Sci. 2019 Jan;1435(1):79-92. doi: 10.1111/nyas.13926. Epub 2018 Jul 13. Ann N Y Acad Sci. 2019. PMID: 30003536 Review.
-
The Wound Microbiome: Modern Approaches to Examining the Role of Microorganisms in Impaired Chronic Wound Healing.Adv Wound Care (New Rochelle). 2014 Jul 1;3(7):502-510. doi: 10.1089/wound.2012.0397. Adv Wound Care (New Rochelle). 2014. PMID: 25032070 Free PMC article. Review.
-
The Wound Microbiome.Cold Spring Harb Perspect Biol. 2023 Jun 1;15(6):a041218. doi: 10.1101/cshperspect.a041218. Cold Spring Harb Perspect Biol. 2023. PMID: 36220302 Free PMC article. Review.
Cited by
-
Chemical Profile, Anti-Microbial and Anti-Inflammaging Activities of Santolina rosmarinifolia L. Essential Oil from Portugal.Antibiotics (Basel). 2023 Jan 15;12(1):179. doi: 10.3390/antibiotics12010179. Antibiotics (Basel). 2023. PMID: 36671380 Free PMC article.
-
A Polyclonal SELEX Aptamer Library Allows Differentiation of Candida albicans, C. auris and C. parapsilosis Cells from Human Dermal Fibroblasts.J Fungi (Basel). 2022 Aug 15;8(8):856. doi: 10.3390/jof8080856. J Fungi (Basel). 2022. PMID: 36012844 Free PMC article.
-
Fluid handling by foam wound dressings: From engineering theory to advanced laboratory performance evaluations.Int Wound J. 2024 Feb;21(2):e14674. doi: 10.1111/iwj.14674. Int Wound J. 2024. PMID: 38353372 Free PMC article. Review.
-
Shockwaves Increase In Vitro Resilience of Rhizopus oryzae Biofilm under Amphotericin B Treatment.Int J Mol Sci. 2022 Aug 17;23(16):9226. doi: 10.3390/ijms23169226. Int J Mol Sci. 2022. PMID: 36012494 Free PMC article.
-
Therapy of infected wounds: overcoming clinical challenges by advanced drug delivery systems.Drug Deliv Transl Res. 2021 Aug;11(4):1545-1567. doi: 10.1007/s13346-021-00932-7. Epub 2021 Feb 20. Drug Deliv Transl Res. 2021. PMID: 33611768 Free PMC article. Review.
References
-
- Wolcott R. Economic aspects of biofilm-based wound care in diabetic foot ulcers. J Wound Care 2015;24:189–190, 192–194 - PubMed
-
- Yarza P, Yilmaz P, Pruesse E, et al. . Uniting the classification of cultured and uncultured bacteria and archaea using 16S rRNA gene sequences. Nat Rev Microbiol 2014;12:635–645 - PubMed
Publication types
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Miscellaneous

