Translation elongation and mRNA stability are coupled through the ribosomal A-site
- PMID: 29997263
- PMCID: PMC6140462
- DOI: 10.1261/rna.066787.118
Translation elongation and mRNA stability are coupled through the ribosomal A-site
Abstract
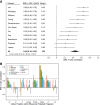
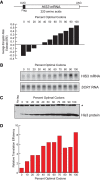
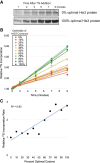
Messenger RNA (mRNA) degradation plays a critical role in regulating transcript levels in eukaryotic cells. Previous work by us and others has shown that codon identity exerts a powerful influence on mRNA stability. In Saccharomyces cerevisiae, studies using a handful of reporter mRNAs show that optimal codons increase translation elongation rate, which in turn increases mRNA stability. However, a direct relationship between elongation rate and mRNA stability has not been established across the entire yeast transcriptome. In addition, there is evidence from work in higher eukaryotes that amino acid identity influences mRNA stability, raising the question as to whether the impact of translation elongation on mRNA decay is at the level of tRNA decoding, amino acid incorporation, or some combination of each. To address these questions, we performed ribosome profiling of wild-type yeast. In good agreement with other studies, our data showed faster codon-specific elongation over optimal codons and faster transcript-level elongation correlating with transcript optimality. At both the codon-level and transcript-level, faster elongation correlated with increased mRNA stability. These findings were reinforced by showing increased translation efficiency and kinetics for a panel of 11 HIS3 reporter mRNAs of increasing codon optimality. While we did observe that elongation measured by ribosome profiling is composed of both amino acid identity and synonymous codon effects, further analyses of these data establish that A-site tRNA decoding rather than other steps of translation elongation is driving mRNA decay in yeast.
Keywords: codon optimality; decoding; mRNA decay; translation elongation.
© 2018 Hanson et al.; Published by Cold Spring Harbor Laboratory Press for the RNA Society.
Figures


References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
