Single-Cell Approach to Influenza-Specific CD8+ T Cell Receptor Repertoires Across Different Age Groups, Tissues, and Following Influenza Virus Infection
- PMID: 29997621
- PMCID: PMC6030351
- DOI: 10.3389/fimmu.2018.01453
Single-Cell Approach to Influenza-Specific CD8+ T Cell Receptor Repertoires Across Different Age Groups, Tissues, and Following Influenza Virus Infection
Abstract
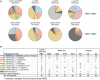
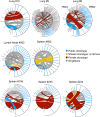
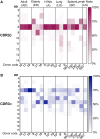
CD8+ T cells recognizing antigenic peptides derived from conserved internal viral proteins confer broad protection against distinct influenza viruses. As memory CD8+ T cells change throughout the human lifetime and across tissue compartments, we investigated how T cell receptor (TCR) composition and diversity relate to memory CD8+ T cells across anatomical sites and immunological phases of human life. We used ex vivo peptide-HLA tetramer magnetic enrichment, single-cell multiplex RT-PCR for both the TCR-alpha (TCRα) and TCR-beta (TCRβ) chains, and new TCRdist and grouping of lymphocyte interactions by paratope hotspots (GLIPH) algorithms to compare TCRs directed against the most prominent human influenza epitope, HLA-A*02:01-M158-66 (A2+M158). We dissected memory TCR repertoires directed toward A2+M158 CD8+ T cells within human tissues and compared them to human peripheral blood of young and elderly adults. Furthermore, we compared these memory CD8+ T cell repertoires to A2+M158 CD8+ TCRs during acute influenza disease in patients hospitalized with avian A/H7N9 virus. Our study provides the first ex vivo comparative analysis of paired antigen-specific TCR-α/β clonotypes across different tissues and peripheral blood across different age groups. We show that human A2+M158 CD8+ T cells can be readily detected in human lungs, spleens, and lymph nodes, and that tissue A2+M158 TCRαβ repertoires reflect A2+M158 TCRαβ clonotypes derived from peripheral blood in healthy adults and influenza-infected patients. A2+M158 TCRαβ repertoires displayed distinct features only in elderly adults, with large private TCRαβ clonotypes replacing the prominent and public TRBV19/TRAV27 TCRs. Our study provides novel findings on influenza-specific TCRαβ repertoires within human tissues, raises the question of how we can prevent the loss of optimal TCRαβ signatures with aging, and provides important insights into the rational design of T cell-mediated vaccines and immunotherapies.
Keywords: CD8+ T cells; T cell receptor repertoire; aging; human T lymphocytes; influenza A virus; tissues.
Figures







Similar articles
-
Perturbed CD8+ T cell immunity across universal influenza epitopes in the elderly.J Leukoc Biol. 2018 Feb;103(2):321-339. doi: 10.1189/jlb.5MA0517-207R. Epub 2017 Dec 29. J Leukoc Biol. 2018. PMID: 28928269
-
Central memory T cells with key TCR repertoires and gene expression profiles dominate influenza CD8+ T cell pools across the human lifespan.Proc Natl Acad Sci U S A. 2025 Jul 29;122(30):e2501167122. doi: 10.1073/pnas.2501167122. Epub 2025 Jul 22. Proc Natl Acad Sci U S A. 2025. PMID: 40694338 Free PMC article.
-
HLA-B*27:05 alters immunodominance hierarchy of universal influenza-specific CD8+ T cells.PLoS Pathog. 2020 Aug 4;16(8):e1008714. doi: 10.1371/journal.ppat.1008714. eCollection 2020 Aug. PLoS Pathog. 2020. PMID: 32750095 Free PMC article.
-
Interrogating the relationship between naïve and immune antiviral T cell repertoires.Curr Opin Virol. 2013 Aug;3(4):447-51. doi: 10.1016/j.coviro.2013.06.011. Epub 2013 Jul 10. Curr Opin Virol. 2013. PMID: 23849601 Free PMC article. Review.
-
Fighting flu: novel CD8+ T-cell targets are required for future influenza vaccines.Clin Transl Immunology. 2024 Feb 14;13(2):e1491. doi: 10.1002/cti2.1491. eCollection 2024. Clin Transl Immunology. 2024. PMID: 38362528 Free PMC article. Review.
Cited by
-
Profiling of T Cell Repertoire in SARS-CoV-2-Infected COVID-19 Patients Between Mild Disease and Pneumonia.J Clin Immunol. 2021 Aug;41(6):1131-1145. doi: 10.1007/s10875-021-01045-z. Epub 2021 May 5. J Clin Immunol. 2021. PMID: 33950324 Free PMC article.
-
Deciphering the TCR Repertoire to Solve the COVID-19 Mystery.Trends Pharmacol Sci. 2020 Aug;41(8):518-530. doi: 10.1016/j.tips.2020.06.001. Epub 2020 Jun 20. Trends Pharmacol Sci. 2020. PMID: 32576386 Free PMC article. Review.
-
HLA class I-associated expansion of TRBV11-2 T cells in multisystem inflammatory syndrome in children.J Clin Invest. 2021 May 17;131(10):e146614. doi: 10.1172/JCI146614. J Clin Invest. 2021. PMID: 33705359 Free PMC article.
-
Lung-Resident Memory CD8+ T Cells in Human Influenza: How Innate Are They?Am J Respir Crit Care Med. 2021 Oct 1;204(7):753-755. doi: 10.1164/rccm.202106-1508ED. Am J Respir Crit Care Med. 2021. PMID: 34352192 Free PMC article. No abstract available.
-
SIV-specific CD8+ T cells are clonotypically distinct across lymphoid and mucosal tissues.J Clin Invest. 2020 Feb 3;130(2):789-798. doi: 10.1172/JCI129161. J Clin Invest. 2020. PMID: 31661461 Free PMC article.
References
-
- Nguyen Thi. H. O, Koutsakos M, Grant EJ, Doherty PC, Kedzierska K. Towards future T cell-mediated influenza vaccines. Infect Dis Transl Med (2016) 2(1):20–9.10.11979/idtm.201601004 - DOI
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

