Dynamics of Dystrophin's Actin-Binding Domain
- PMID: 30007583
- PMCID: PMC6084637
- DOI: 10.1016/j.bpj.2018.05.039
Dynamics of Dystrophin's Actin-Binding Domain
Abstract
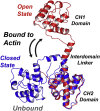
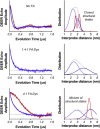
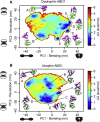
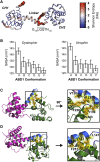
We have used pulsed electron paramagnetic resonance, calorimetry, and molecular dynamics simulations to examine the structural mechanism of binding for dystrophin's N-terminal actin-binding domain (ABD1) and compare it to utrophin's ABD1. Like other members of the spectrin superfamily, dystrophin's ABD1 consists of two calponin-homology (CH) domains, CH1 and CH2. Several mutations within dystrophin's ABD1 are associated with the development of severe degenerative muscle disorders Duchenne and Becker muscular dystrophies, highlighting the importance of understanding its structural biology. To investigate structural changes within dystrophin ABD1 upon binding to actin, we labeled the protein with spin probes and measured changes in inter-CH domain distance using double-electron electron resonance. Previous studies on the homologous protein utrophin showed that actin binding induces a complete structural opening of the CH domains, resulting in a highly ordered ABD1-actin complex. In this study, double-electron electron resonance shows that dystrophin ABD1 also undergoes a conformational opening upon binding F-actin, but this change is less complete and significantly more structurally disordered than observed for utrophin. Using molecular dynamics simulations, we identified a hinge in the linker region between the two CH domains that grants conformational flexibility to ABD1. The conformational dynamics of both dystrophin's and utrophin's ABD1 showed that compact conformations driven by hydrophobic interactions are preferred and that extended conformations are energetically accessible through a flat free-energy surface. Considering that the binding free energy of ABD1 to actin is on the order of 6-7 kcal/mole, our data are compatible with a mechanism in which binding to actin is largely dictated by specific interactions with CH1, but fine tuning of the binding affinity is achieved by the overlap between conformational ensembles of ABD1 free and bound to actin.
Copyright © 2018 Biophysical Society. Published by Elsevier Inc. All rights reserved.
Figures
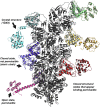
References
-
- Koenig M., Monaco A.P., Kunkel L.M. The complete sequence of dystrophin predicts a rod-shaped cytoskeletal protein. Cell. 1988;53:219–228. - PubMed
-
- Ervasti J.M. Dystrophin, its interactions with other proteins, and implications for muscular dystrophy. Biochim. Biophys. Acta. 2007;1772:108–117. - PubMed
-
- Turner P.R., Westwood T., Steinhardt R.A. Increased protein degradation results from elevated free calcium levels found in muscle from mdx mice. Nature. 1988;335:735–738. - PubMed
-
- Rief M., Pascual J., Gaub H.E. Single molecule force spectroscopy of spectrin repeats: low unfolding forces in helix bundles. J. Mol. Biol. 1999;286:553–561. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

