A Database of Predicted Binding Sites for Cholesterol on Membrane Proteins, Deep in the Membrane
- PMID: 30007584
- PMCID: PMC6084638
- DOI: 10.1016/j.bpj.2018.06.022
A Database of Predicted Binding Sites for Cholesterol on Membrane Proteins, Deep in the Membrane
Abstract
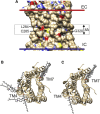
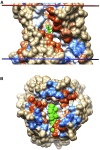
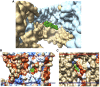
The outer membranes of animal cells contain high concentrations of cholesterol, of which a small proportion is located deep within the hydrophobic core of the membrane. An automated docking procedure is described that allows the characterization of binding sites for these deep cholesterol molecules on the membrane-spanning surfaces of membrane proteins and in protein cavities or pores, driven by hydrogen bond formation. A database of this class of predicted binding site is described, covering 397 high-resolution structures. The database includes sites on the transmembrane surfaces of many G-protein coupled receptors; within the fenestrations of two-pore K+ channels and ATP-gated P2X3 channels; in the central cavities of a number of transporters, including Glut1, Glut5, and P-glycoprotein; and in deep clefts in mitochondrial complexes III and IV.
Copyright © 2018 Biophysical Society. Published by Elsevier Inc. All rights reserved.
Figures


Similar articles
-
G protein coupled receptor interactions with cholesterol deep in the membrane.Biochim Biophys Acta Biomembr. 2017 Feb;1859(2):268-281. doi: 10.1016/j.bbamem.2016.12.001. Epub 2016 Dec 3. Biochim Biophys Acta Biomembr. 2017. PMID: 27919726
-
Interfacial Binding Sites for Cholesterol on Kir, Kv, K2P, and Related Potassium Channels.Biophys J. 2020 Jul 7;119(1):35-47. doi: 10.1016/j.bpj.2020.05.028. Epub 2020 Jun 4. Biophys J. 2020. PMID: 32553129 Free PMC article.
-
Interfacial Binding Sites for Cholesterol on G Protein-Coupled Receptors.Biophys J. 2019 May 7;116(9):1586-1597. doi: 10.1016/j.bpj.2019.03.025. Epub 2019 Apr 2. Biophys J. 2019. PMID: 31010663 Free PMC article.
-
K(ATP) channels "vingt ans après": ATG to PDB to Mechanism.J Mol Cell Cardiol. 2005 Jul;39(1):79-98. doi: 10.1016/j.yjmcc.2004.12.004. Epub 2005 Feb 19. J Mol Cell Cardiol. 2005. PMID: 15978905 Review.
-
Mechanism of StAR's regulation of mitochondrial cholesterol import.Mol Cell Endocrinol. 2007 Feb;265-266:46-50. doi: 10.1016/j.mce.2006.12.002. Epub 2007 Jan 5. Mol Cell Endocrinol. 2007. PMID: 17207924 Review.
Cited by
-
Databases of ligand-binding pockets and protein-ligand interactions.Comput Struct Biotechnol J. 2024 Mar 24;23:1320-1338. doi: 10.1016/j.csbj.2024.03.015. eCollection 2024 Dec. Comput Struct Biotechnol J. 2024. PMID: 38585646 Free PMC article. Review.
-
Improved Highly Mobile Membrane Mimetic Model for Investigating Protein-Cholesterol Interactions.J Chem Inf Model. 2024 Jun 24;64(12):4822-4834. doi: 10.1021/acs.jcim.4c00619. Epub 2024 Jun 6. J Chem Inf Model. 2024. PMID: 38844760 Free PMC article.
-
Interfacial Binding Sites for Cholesterol on TRP Ion Channels.Biophys J. 2019 Nov 19;117(10):2020-2033. doi: 10.1016/j.bpj.2019.10.011. Epub 2019 Oct 18. Biophys J. 2019. PMID: 31672270 Free PMC article.
-
The energetics of protein-lipid interactions as viewed by molecular simulations.Biochem Soc Trans. 2020 Feb 28;48(1):25-37. doi: 10.1042/BST20190149. Biochem Soc Trans. 2020. PMID: 31872229 Free PMC article. Review.
-
Interfacial binding sites for cholesterol on GABAA receptors and competition with neurosteroids.Biophys J. 2021 Jul 6;120(13):2710-2722. doi: 10.1016/j.bpj.2021.05.009. Epub 2021 May 20. Biophys J. 2021. PMID: 34022235 Free PMC article.
References
-
- Genheden S., Essex J.W., Lee A.G. G protein coupled receptor interactions with cholesterol deep in the membrane. Biochim. Biophys. Acta. 2017;1859:268–281. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Miscellaneous

