Formal description of sequence-based voucherless Fungi: promises and pitfalls, and how to resolve them
- PMID: 30018876
- PMCID: PMC6048566
- DOI: 10.5598/imafungus.2018.09.01.09
Formal description of sequence-based voucherless Fungi: promises and pitfalls, and how to resolve them
Abstract
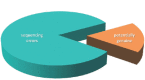
There is urgent need for a formal nomenclature of sequence-based, voucherless Fungi, given that environmental sequencing has accumulated more than one billion fungal ITS reads in the Sequence Read Archive, about 1,000 times as many as fungal ITS sequences in GenBank. These unnamed Fungi could help to bridge the gap between 115,000 to 140,000 currently accepted and 2.2 to 3.8 million predicted species, a gap that cannot realistically be filled using specimen or culture-based inventories. The Code never aimed at placing restrictions on the nature of characters chosen for taxonomy, and the requirement for physical types is now becoming a constraint on the advancement of science. We elaborate on the promises and pitfalls of sequence-based nomenclature and provide potential solutions to major concerns of the mycological community. Types of sequence-based taxa, which by default lack a physical specimen or culture, could be designated in four alternative ways: (1) the underlying sample ('bag' type), (2) the DNA extract, (3) fluorescent in situ hybridization (FISH), or (4) the type sequence itself. Only (4) would require changes to the Code and the latter would be the most straightforward approach, complying with three of the five principal functions of types better than physical specimens. A fifth way, representation of the sequence in an illustration, has been ruled as unacceptable in the Code. Potential flaws in sequence data are analogous to flaws in physical types, and artifacts are manageable if a stringent analytical approach is applied. Conceptual errors such as homoplasy, intragenomic variation, gene duplication, hybridization, and horizontal gene transfer, apply to all molecular approaches and cannot be used as a specific argument against sequence-based nomenclature. The potential impact of these phenomena is manageable, as phylogenetic species delimitation has worked satisfactorily in Fungi. The most serious shortcoming of sequence-based nomenclature is the likelihood of parallel classifications, either by describing taxa that already have names based on physical types, or by using different markers to delimit species within the same lineage. The probability of inadvertently establishing sequence-based species that have names available is between 20.4 % and 1.5 % depending on the number of globally predicted fungal species. This compares favourably to a historical error rate of about 30 % based on physical types, and this rate could be reduced to practically zero by adding specific provisions to this approach in the Code. To avoid parallel classifications based on different markers, sequence-based nomenclature should be limited to a single marker, preferably the fungal ITS barcoding marker; this is possible since sequence-based nomenclature does not aim at accurate species delimitation but at naming lineages to generate a reference database, independent of whether these lineages represent species, closely related species complexes, or infraspecies. We argue that clustering methods are inappropriate for sequence-based nomenclature; this approach must instead use phylogenetic methods based on multiple alignments, combined with quantitative species recognition methods. We outline strategies to obtain higher-level phylogenies for ITS-based, voucherless species, including phylogenetic binning, 'hijacking' species delimitation methods, and temporal banding. We conclude that voucherless, sequence-based nomenclature is not a threat to specimen and culture-based fungal taxonomy, but a complementary approach capable of substantially closing the gap between known and predicted fungal diversity, an approach that requires careful work and high skill levels.
Keywords: IMC11; biodiversity; ecologically cryptic Fungi; environmental sequencing; evolutionary placement algorithm; high throughput sequencing; internal transcribed spacer; molecular barcoding; molecular sequence data; next generation sequencing; nomenclature; typification.
Figures


References
-
- Aguileta G, Marthey S, Chiapello H, Lebrun MH, Rodolphe F, Fournier E, Gendrault-Jacquemard A, Giraud T. (2008) Assessing the performance of single-copy genes for recovering robust phylogenies. Systematic Biology 57: 613–627. - PubMed
-
- Arnold AE, Henk DA, Eells RL, Lutzoni F, Vilgalys R. (2007) Diversity and phylogenetic affinities of foliar fungal endophytes in loblolly pine inferred by culturing and environmental PCR. Mycologia 99: 185–206. - PubMed
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous
