Uncovering the hidden half of plants using new advances in root phenotyping
- PMID: 30031961
- PMCID: PMC6378649
- DOI: 10.1016/j.copbio.2018.06.002
Uncovering the hidden half of plants using new advances in root phenotyping
Abstract
Major increases in crop yield are required to keep pace with population growth and climate change. Improvements to the architecture of crop roots promise to deliver increases in water and nutrient use efficiency but profiling the root phenome (i.e. its structure and function) represents a major bottleneck. We describe how advances in imaging and sensor technologies are making root phenomic studies possible. However, methodological advances in acquisition, handling and processing of the resulting 'big-data' is becoming increasingly important. Advances in automated image analysis approaches such as Deep Learning promise to transform the root phenotyping landscape. Collectively, these innovations are helping drive the selection of the next-generation of crops to deliver real world impact for ongoing global food security efforts.
Copyright © 2018 The Authors. Published by Elsevier Ltd.. All rights reserved.
Figures

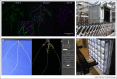


References
-
- Lynch J.P. Roots of the second green revolution. Aust J Bot. 2007;55:493–512.
-
- Topp C.N., Iyer-Pascuzzi A.S., Anderson J.T., Lee C.-R., Zurek P.R., Symonova O., Zheng Y., Bucksch A., Mileyko Y., Galkovskyi T. 3D phenotyping and quantitative trait locus mapping identify core regions of the rice genome controlling root architecture. Proc Natl Acad Sci U S A. 2013;110:E1695–E1704. - PMC - PubMed
-
The authors developed a 3D phenotyping platform based on transparent media and used it to identify QTL for root architecture in rice.
-
- Wang Q., Komarov S., Mathews A.J., Li K., Topp C., O’Sullivan J.A., Y-C Tai. 2014. Combined 3D PET and optical projection tomography techniques for plant root phenotyping. ArXiv150100242 Phys Q-Bio.
-
- Rellán-Álvarez R., Lobet G., Lindner H., Pradier P.-L., Sebastian J., Yee M.-C., Geng Y., Trontin C., LaRue T., Schrager-Lavelle A. GLO-Roots: an imaging platform enabling multidimensional characterization of soil-grown root systems. eLife. 2015;4:e07597. - PMC - PubMed
-
The authors developed an innovative luciferase reporter based approach to image transgenic roots in rhizotrons. The GLO-ROOTS approach provides novel opportunities to study the impact of soil properties on root gene expression.
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

