Enzymes and Mechanisms Employed by Tailed Bacteriophages to Breach the Bacterial Cell Barriers
- PMID: 30060520
- PMCID: PMC6116005
- DOI: 10.3390/v10080396
Enzymes and Mechanisms Employed by Tailed Bacteriophages to Breach the Bacterial Cell Barriers
Abstract
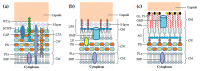
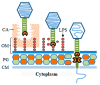
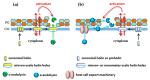
Monoderm bacteria possess a cell envelope made of a cytoplasmic membrane and a cell wall, whereas diderm bacteria have and extra lipid layer, the outer membrane, covering the cell wall. Both cell types can also produce extracellular protective coats composed of polymeric substances like, for example, polysaccharidic capsules. Many of these structures form a tight physical barrier impenetrable by phage virus particles. Tailed phages evolved strategies/functions to overcome the different layers of the bacterial cell envelope, first to deliver the genetic material to the host cell cytoplasm for virus multiplication, and then to release the virion offspring at the end of the reproductive cycle. There is however a major difference between these two crucial steps of the phage infection cycle: virus entry cannot compromise cell viability, whereas effective virion progeny release requires host cell lysis. Here we present an overview of the viral structures, key protein players and mechanisms underlying phage DNA entry to bacteria, and then escape of the newly-formed virus particles from infected hosts. Understanding the biological context and mode of action of the phage-derived enzymes that compromise the bacterial cell envelope may provide valuable information for their application as antimicrobials.
Keywords: LysB; bacterial cell envelope; cell wall; depolymerase; endolysin; holin; lysin; peptidoglycan; proton motive force; spanin.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures
Similar articles
-
Phage Lysis: Multiple Genes for Multiple Barriers.Adv Virus Res. 2019;103:33-70. doi: 10.1016/bs.aivir.2018.09.003. Epub 2018 Nov 28. Adv Virus Res. 2019. PMID: 30635077 Free PMC article. Review.
-
Phage lysis: three steps, three choices, one outcome.J Microbiol. 2014 Mar;52(3):243-58. doi: 10.1007/s12275-014-4087-z. Epub 2014 Mar 1. J Microbiol. 2014. PMID: 24585055 Free PMC article. Review.
-
Bacteriophage and peptidoglycan degrading enzymes with antimicrobial applications.Recent Pat Biotechnol. 2007;1(2):113-22. doi: 10.2174/187220807780809463. Recent Pat Biotechnol. 2007. PMID: 19075835 Review.
-
Endolysin Regulation in Phage Mu Lysis.mBio. 2022 Jun 28;13(3):e0081322. doi: 10.1128/mbio.00813-22. Epub 2022 Apr 26. mBio. 2022. PMID: 35471081 Free PMC article.
-
Enterococcal Bacteriophages and Genome Defense.2014 Feb 11. In: Gilmore MS, Clewell DB, Ike Y, Shankar N, editors. Enterococci: From Commensals to Leading Causes of Drug Resistant Infection [Internet]. Boston: Massachusetts Eye and Ear Infirmary; 2014–. 2014 Feb 11. In: Gilmore MS, Clewell DB, Ike Y, Shankar N, editors. Enterococci: From Commensals to Leading Causes of Drug Resistant Infection [Internet]. Boston: Massachusetts Eye and Ear Infirmary; 2014–. PMID: 24649501 Free Books & Documents. Review.
Cited by
-
Bacteriophage-mediated approaches for biofilm control.Front Cell Infect Microbiol. 2024 Oct 7;14:1428637. doi: 10.3389/fcimb.2024.1428637. eCollection 2024. Front Cell Infect Microbiol. 2024. PMID: 39435185 Free PMC article. Review.
-
Analysis of a Novel Bacteriophage vB_AchrS_AchV4 Highlights the Diversity of Achromobacter Viruses.Viruses. 2021 Feb 27;13(3):374. doi: 10.3390/v13030374. Viruses. 2021. PMID: 33673419 Free PMC article.
-
Tall tails: cryo-electron microscopy of phage tail DNA ejection conduits.Biochem Soc Trans. 2022 Feb 28;50(1):459-22W. doi: 10.1042/BST20210799. Biochem Soc Trans. 2022. PMID: 35129586 Free PMC article.
-
Are Bordetella bronchiseptica Siphoviruses (Genus Vojvodinavirus) Appropriate for Phage Therapy-Bacterial Allies or Foes?Viruses. 2021 Aug 31;13(9):1732. doi: 10.3390/v13091732. Viruses. 2021. PMID: 34578315 Free PMC article.
-
|Isolation and characterization of novel bacteriophages as a potential therapeutic option for Escherichia coli urinary tract infections.Appl Microbiol Biotechnol. 2021 Jul;105(13):5617-5629. doi: 10.1007/s00253-021-11432-6. Epub 2021 Jul 12. Appl Microbiol Biotechnol. 2021. PMID: 34254156 Free PMC article.
References
-
- Carter J., Saunders V. Virology: Principles and Applications. 2nd ed. Wiley & Sons Inc.; New York, NY, USA: 2013.
-
- São-José C., Nascimento J., Parreira R., Santos M. Release of progeny phages from infected cells. In: McGrath S., van Sinderen D., editors. Bacteriophage: Genetics and Molecular Biology. Caister Academic Press; Norfolk, UK: 2007. pp. 307–333.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

