Culture- and metagenomics-enabled analyses of the Methanosphaera genus reveals their monophyletic origin and differentiation according to genome size
- PMID: 30068938
- PMCID: PMC6246560
- DOI: 10.1038/s41396-018-0225-7
Culture- and metagenomics-enabled analyses of the Methanosphaera genus reveals their monophyletic origin and differentiation according to genome size
Abstract
The genus Methanosphaera is a well-recognized but poorly characterized member of the mammalian gut microbiome, and distinctive from Methanobrevibacter smithii for its ability to induce a pro-inflammatory response in humans. Here we have used a combination of culture- and metagenomics-based approaches to expand the representation and information for the genus, which has supported the examination of their phylogeny and physiological capacity. Novel isolates of the genus Methanosphaera were recovered from bovine rumen digesta and human stool, with the bovine isolate remarkable for its large genome size relative to other Methanosphaera isolates from monogastric hosts. To substantiate this observation, we then recovered seven high-quality Methanosphaera-affiliated population genomes from ruminant and human gut metagenomic datasets. Our analyses confirm a monophyletic origin of Methanosphaera spp. and that the colonization of monogastric and ruminant hosts favors representatives of the genus with different genome sizes, reflecting differences in the genome content needed to persist in these different habitats.
Conflict of interest statement
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
E.C.H. planned and performed research and helped write the paper; D.H.P., J.G.V., C.P.R., C.S.M., S.E.D. and P.Ó.C. assisted with research and helped write the paper. P.R.G. and J.G.M. provided samples enabling
Figures

 , 1.3
, 1.3  , 1.8
, 1.8  , 2.6
, 2.6  , 3.5
, 3.5  and 4.6 mM) or in the instance of (d), ethanol (0
and 4.6 mM) or in the instance of (d), ethanol (0  , 3
, 3  , 10
, 10  , 30
, 30  , 50
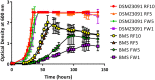
, 50  and 171 mM). The growth of the individual cultures is plotted until the time point at which maximal OD600 was measured. Individual values represent the mean (±SD) produced from triplicate cultures
and 171 mM). The growth of the individual cultures is plotted until the time point at which maximal OD600 was measured. Individual values represent the mean (±SD) produced from triplicate cultures


References
-
- Brusa T, Canzi E, Allievi L, Puppo E, Ferrari A. Methanogens in the human intestinal tract and oral cavity. Curr Microbiol. 1993;27:261–5. doi: 10.1007/BF01575989. - DOI
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

