Co-regulatory networks of human serum proteins link genetics to disease
- PMID: 30072576
- PMCID: PMC6190714
- DOI: 10.1126/science.aaq1327
Co-regulatory networks of human serum proteins link genetics to disease
Abstract
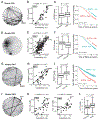
Proteins circulating in the blood are critical for age-related disease processes; however, the serum proteome has remained largely unexplored. To this end, 4137 proteins covering most predicted extracellular proteins were measured in the serum of 5457 Icelanders over 65 years of age. Pairwise correlation between proteins as they varied across individuals revealed 27 different network modules of serum proteins, many of which were associated with cardiovascular and metabolic disease states, as well as overall survival. The protein modules were controlled by cis- and trans-acting genetic variants, which in many cases were also associated with complex disease. This revealed co-regulated groups of circulating proteins that incorporated regulatory control between tissues and demonstrated close relationships to past, current, and future disease states.
Copyright © 2018 The Authors, some rights reserved; exclusive licensee American Association for the Advancement of Science. No claim to original U.S. Government Works.
Conflict of interest statement
Figures
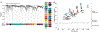


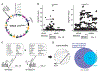
Comment in
-
Blood Is a Window into Health and Disease.Clin Chem. 2019 Oct;65(10):1204-1206. doi: 10.1373/clinchem.2018.299065. Epub 2019 Jun 6. Clin Chem. 2019. PMID: 31171530 No abstract available.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Miscellaneous

