The voltage-gated sodium channel pore exhibits conformational flexibility during slow inactivation
- PMID: 30082431
- PMCID: PMC6122925
- DOI: 10.1085/jgp.201812118
The voltage-gated sodium channel pore exhibits conformational flexibility during slow inactivation
Abstract
Slow inactivation in voltage-gated sodium channels (NaVs) directly regulates the excitability of neurons, cardiac myocytes, and skeletal muscles. Although NaV slow inactivation appears to be conserved across phylogenies-from bacteria to humans-the structural basis for this mechanism remains unclear. Here, using site-directed labeling and EPR spectroscopic measurements of membrane-reconstituted prokaryotic NaV homologues, we characterize the conformational dynamics of the selectivity filter region in the conductive and slow-inactivated states to determine the molecular events underlying NaV gating. Our findings reveal profound conformational flexibility of the pore in the slow-inactivated state. We find that the P1 and P2 pore helices undergo opposing movements with respect to the pore axis. These movements result in changes in volume of both the central and intersubunit cavities, which form pathways for lipophilic drugs that modulate slow inactivation. Our findings therefore provide novel insight into the molecular basis for state-dependent effects of lipophilic drugs on channel function.
© 2018 Chatterjee et al.
Figures


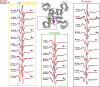




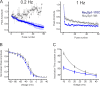

Comment in
-
Progress in understanding slow inactivation speeds up.J Gen Physiol. 2018 Sep 3;150(9):1235-1238. doi: 10.1085/jgp.201812149. Epub 2018 Aug 16. J Gen Physiol. 2018. PMID: 30115663 Free PMC article.
References
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

