The Dysregulation of the DLK1- MEG3 Locus in Islets From Patients With Type 2 Diabetes Is Mimicked by Targeted Epimutation of Its Promoter With TALE-DNMT Constructs
- PMID: 30084829
- PMCID: PMC6110314
- DOI: 10.2337/db17-0682
The Dysregulation of the DLK1- MEG3 Locus in Islets From Patients With Type 2 Diabetes Is Mimicked by Targeted Epimutation of Its Promoter With TALE-DNMT Constructs
Abstract
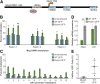
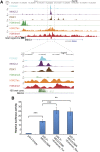
Type 2 diabetes mellitus (T2DM) is characterized by the inability of the insulin-producing β-cells to overcome insulin resistance. We previously identified an imprinted region on chromosome 14, the DLK1-MEG3 locus, as being downregulated in islets from humans with T2DM. In this study, using targeted epigenetic modifiers, we prove that increased methylation at the promoter of Meg3 in mouse βTC6 β-cells results in decreased transcription of the maternal transcripts associated with this locus. As a result, the sensitivity of β-cells to cytokine-mediated oxidative stress was increased. Additionally, we demonstrate that an evolutionarily conserved intronic region at the MEG3 locus can function as an enhancer in βTC6 β-cells. Using circular chromosome conformation capture followed by high-throughput sequencing, we demonstrate that the promoter of MEG3 physically interacts with this novel enhancer and other putative regulatory elements in this imprinted region in human islets. Remarkably, this enhancer is bound in an allele-specific manner by the transcription factors FOXA2, PDX1, and NKX2.2. Overall, these data suggest that the intronic MEG3 enhancer plays an important role in the regulation of allele-specific expression at the imprinted DLK1-MEG3 locus in human β-cells, which in turn impacts the sensitivity of β-cells to cytokine-mediated oxidative stress.
© 2018 by the American Diabetes Association.
Figures


Similar articles
-
Epigenetic regulation of the DLK1-MEG3 microRNA cluster in human type 2 diabetic islets.Cell Metab. 2014 Jan 7;19(1):135-45. doi: 10.1016/j.cmet.2013.11.016. Epub 2013 Dec 26. Cell Metab. 2014. PMID: 24374217 Free PMC article.
-
Analysis of the Paternally-Imprinted DLK1-MEG3 and IGF2-H19 Tandem Gene Loci in NT2 Embryonal Carcinoma Cells Identifies DLK1 as a Potential Therapeutic Target.Stem Cell Rev Rep. 2018 Dec;14(6):823-836. doi: 10.1007/s12015-018-9838-5. Stem Cell Rev Rep. 2018. PMID: 29980981
-
Dlk1-Dio3 locus-derived lncRNAs perpetuate postmitotic motor neuron cell fate and subtype identity.Elife. 2018 Oct 12;7:e38080. doi: 10.7554/eLife.38080. Elife. 2018. PMID: 30311912 Free PMC article.
-
DLK1-DIO3 imprinted cluster in induced pluripotency: landscape in the mist.Cell Mol Life Sci. 2014 Nov;71(22):4421-30. doi: 10.1007/s00018-014-1698-9. Epub 2014 Aug 7. Cell Mol Life Sci. 2014. PMID: 25098353 Free PMC article. Review.
-
MEG3 noncoding RNA: a tumor suppressor.J Mol Endocrinol. 2012 Apr 2;48(3):R45-53. doi: 10.1530/JME-12-0008. Print 2012 Jun. J Mol Endocrinol. 2012. PMID: 22393162 Free PMC article. Review.
Cited by
-
Bioinformatics Analysis of Next Generation Sequencing Data Identifies Molecular Biomarkers Associated With Type 2 Diabetes Mellitus.Clin Med Insights Endocrinol Diabetes. 2023 Feb 20;16:11795514231155635. doi: 10.1177/11795514231155635. eCollection 2023. Clin Med Insights Endocrinol Diabetes. 2023. PMID: 36844983 Free PMC article.
-
Genetic regulation of RNA splicing in human pancreatic islets.Genome Biol. 2022 Sep 15;23(1):196. doi: 10.1186/s13059-022-02757-0. Genome Biol. 2022. PMID: 36109769 Free PMC article.
-
Enzyme-free targeted DNA demethylation using CRISPR-dCas9-based steric hindrance to identify DNA methylation marks causal to altered gene expression.Nat Protoc. 2022 Dec;17(12):2840-2881. doi: 10.1038/s41596-022-00741-3. Epub 2022 Oct 7. Nat Protoc. 2022. PMID: 36207463 Review.
-
Placental maternally expressed gene 3 differentially methylated region methylation profile is associated with maternal glucose concentration and newborn birthweight.J Diabetes Investig. 2021 Jun;12(6):1074-1082. doi: 10.1111/jdi.13432. Epub 2020 Nov 4. J Diabetes Investig. 2021. PMID: 33090678 Free PMC article.
-
Non-Coding RNA in Pancreas and β-Cell Development.Noncoding RNA. 2018 Dec 13;4(4):41. doi: 10.3390/ncrna4040041. Noncoding RNA. 2018. PMID: 30551650 Free PMC article. Review.
References
-
- da Rocha ST, Edwards CA, Ito M, Ogata T, Ferguson-Smith AC. Genomic imprinting at the mammalian Dlk1-Dio3 domain. Trends Genet 2008;24:306–316 - PubMed
-
- You L, Wang N, Yin D, et al. . Downregulation of long noncoding RNA Meg3 affects insulin synthesis and secretion in mouse pancreatic beta cells. J Cell Physiol 2016;231:852–862 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials

