UM171 Enhances Lentiviral Gene Transfer and Recovery of Primitive Human Hematopoietic Cells
- PMID: 30101153
- PMCID: PMC6077133
- DOI: 10.1016/j.omtm.2018.06.009
UM171 Enhances Lentiviral Gene Transfer and Recovery of Primitive Human Hematopoietic Cells
Abstract
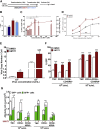
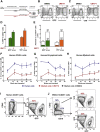
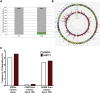
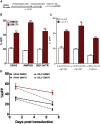
Enhanced gene transfer efficiencies and higher yields of transplantable transduced human hematopoietic stem cells are continuing goals for improving clinical protocols that use stemcell-based gene therapies. Here, we examined the effect of the HSC agonist UM171 on these endpoints in both in vitro and in vivo systems. Using a 22-hr transduction protocol, we found that UM171 significantly enhances both the lentivirus-mediated transduction and yield of CD34+ and CD34+CD45RA- hematopoietic cells from human cord blood to give a 6-fold overall higher recovery of transduced hematopoietic stem cells, including cells with long-term lympho-myeloid repopulating activity in immunodeficient mice. The ability of UM171 to enhance gene transfer to primitive cord blood hematopoietic cells extended to multiple lentiviral pseudotypes, gamma retroviruses, and non-integrating lentiviruses and to adult bone marrow cells. UM171, thus, provides an interesting reagent for improving the ex vivo production of gene-modified cells and for reducing requirements of virus for a broad range of applications.
Keywords: gene transfer; hematopoietic stem cells; lentivirus.
Figures
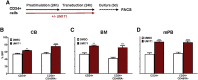
Similar articles
-
High levels of transgene expression following transduction of long-term NOD/SCID-repopulating human cells with a modified lentiviral vector.Stem Cells. 2001;19(3):247-59. doi: 10.1634/stemcells.19-3-247. Stem Cells. 2001. PMID: 11359950
-
Hematopoietic stem cell transplantation using single UM171-expanded cord blood: a single-arm, phase 1-2 safety and feasibility study.Lancet Haematol. 2020 Feb;7(2):e134-e145. doi: 10.1016/S2352-3026(19)30202-9. Epub 2019 Nov 6. Lancet Haematol. 2020. PMID: 31704264 Clinical Trial.
-
Molecular evidence of lentiviral vector-mediated gene transfer into human self-renewing, multi-potent, long-term NOD/SCID repopulating hematopoietic cells.Mol Ther. 2002 Nov;6(5):615-26. Mol Ther. 2002. PMID: 12409260
-
Human cord blood CD34+CD38- cell transduction via lentivirus-based gene transfer vectors.Hum Gene Ther. 1999 Jun 10;10(9):1479-89. doi: 10.1089/10430349950017815. Hum Gene Ther. 1999. PMID: 10395373
-
Role of the mammalian target of rapamycin pathway in lentiviral vector transduction of hematopoietic stem cells.Curr Opin Hematol. 2015 Jul;22(4):302-8. doi: 10.1097/MOH.0000000000000150. Curr Opin Hematol. 2015. PMID: 26049750 Free PMC article. Review.
Cited by
-
Pro-inflammatory macrophages suppress HIV replication in humanized mice and ex vivo co-cultures.Front Immunol. 2024 Nov 7;15:1439328. doi: 10.3389/fimmu.2024.1439328. eCollection 2024. Front Immunol. 2024. PMID: 39575258 Free PMC article.
-
Expanded circulating hematopoietic stem/progenitor cells as novel cell source for the treatment of TCIRG1 osteopetrosis.Haematologica. 2021 Jan 1;106(1):74-86. doi: 10.3324/haematol.2019.238261. Haematologica. 2021. PMID: 31949009 Free PMC article.
-
Advances in hematopoietic stem cells ex vivo expansion associated with bone marrow niche.Ann Hematol. 2024 Dec;103(12):5035-5057. doi: 10.1007/s00277-024-05773-1. Epub 2024 Apr 30. Ann Hematol. 2024. PMID: 38684510 Review.
-
Development of β-globin gene correction in human hematopoietic stem cells as a potential durable treatment for sickle cell disease.Sci Transl Med. 2021 Jun 16;13(598):eabf2444. doi: 10.1126/scitranslmed.abf2444. Sci Transl Med. 2021. PMID: 34135108 Free PMC article.
-
Mouse models in hematopoietic stem cell gene therapy and genome editing.Biochem Pharmacol. 2020 Apr;174:113692. doi: 10.1016/j.bcp.2019.113692. Epub 2019 Nov 6. Biochem Pharmacol. 2020. PMID: 31705854 Free PMC article. Review.
References
-
- Cartier N., Hacein-Bey-Abina S., Bartholomae C.C., Veres G., Schmidt M., Kutschera I., Vidaud M., Abel U., Dal-Cortivo L., Caccavelli L. Hematopoietic stem cell gene therapy with a lentiviral vector in X-linked adrenoleukodystrophy. Science. 2009;326:818–823. - PubMed
LinkOut - more resources
Full Text Sources
Other Literature Sources

