Tousled-like kinases stabilize replication forks and show synthetic lethality with checkpoint and PARP inhibitors
- PMID: 30101194
- PMCID: PMC6082654
- DOI: 10.1126/sciadv.aat4985
Tousled-like kinases stabilize replication forks and show synthetic lethality with checkpoint and PARP inhibitors
Abstract
DNA sequence and epigenetic information embedded in chromatin must be faithfully duplicated and transmitted to daughter cells during cell division. However, how chromatin assembly and DNA replication are integrated remains unclear. We examined the contribution of the Tousled-like kinases 1 and 2 (TLK1/TLK2) to chromatin assembly and maintenance of replication fork integrity. We show that TLK activity is required for DNA replication and replication-coupled nucleosome assembly and that lack of TLK activity leads to replication fork stalling and the accumulation of single-stranded DNA, a phenotype distinct from ASF1 depletion. Consistent with these results, sustained TLK depletion gives rise to replication-dependent DNA damage and p53-dependent cell cycle arrest in G1. We find that deficient replication-coupled de novo nucleosome assembly renders replication forks unstable and highly dependent on the ATR and CHK1 checkpoint kinases, as well as poly(adenosine 5'-diphosphate-ribose) polymerase (PARP) activity, to avoid collapse. Human cancer data revealed frequent up-regulation of TLK genes and an association with poor patient outcome in multiple types of cancer, and depletion of TLK activity leads to increased replication stress and DNA damage in a panel of cancer cells. Our results reveal a critical role for TLKs in chromatin replication and suppression of replication stress and identify a synergistic lethal relationship with checkpoint signaling and PARP that could be exploited in treatment of a broad range of cancers.
Figures


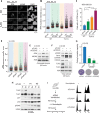

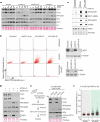

Similar articles
-
Suppression of Tousled-like kinase activity after DNA damage or replication block requires ATM, NBS1 and Chk1.Oncogene. 2003 Sep 4;22(38):5927-37. doi: 10.1038/sj.onc.1206691. Oncogene. 2003. PMID: 12955071
-
Identification of human Asf1 chromatin assembly factors as substrates of Tousled-like kinases.Curr Biol. 2001 Jul 10;11(13):1068-73. doi: 10.1016/s0960-9822(01)00298-6. Curr Biol. 2001. PMID: 11470414
-
Human Tousled like kinases are targeted by an ATM- and Chk1-dependent DNA damage checkpoint.EMBO J. 2003 Apr 1;22(7):1676-87. doi: 10.1093/emboj/cdg151. EMBO J. 2003. PMID: 12660173 Free PMC article.
-
The synthetic lethality of targeting cell cycle checkpoints and PARPs in cancer treatment.J Hematol Oncol. 2022 Oct 17;15(1):147. doi: 10.1186/s13045-022-01360-x. J Hematol Oncol. 2022. PMID: 36253861 Free PMC article. Review.
-
The Tousled-like kinases regulate genome and epigenome stability: implications in development and disease.Cell Mol Life Sci. 2019 Oct;76(19):3827-3841. doi: 10.1007/s00018-019-03208-z. Epub 2019 Jul 13. Cell Mol Life Sci. 2019. PMID: 31302748 Free PMC article. Review.
Cited by
-
Identification of DNA damage response-related genes as biomarkers for castration-resistant prostate cancer.Sci Rep. 2023 Nov 10;13(1):19602. doi: 10.1038/s41598-023-46651-6. Sci Rep. 2023. PMID: 37950047 Free PMC article.
-
Advances in synthetic lethality for cancer therapy: cellular mechanism and clinical translation.J Hematol Oncol. 2020 Sep 3;13(1):118. doi: 10.1186/s13045-020-00956-5. J Hematol Oncol. 2020. PMID: 32883316 Free PMC article. Review.
-
Exploiting the DNA Damage Response for Prostate Cancer Therapy.Cancers (Basel). 2023 Dec 23;16(1):83. doi: 10.3390/cancers16010083. Cancers (Basel). 2023. PMID: 38201511 Free PMC article. Review.
-
Identification of modulators of the ALT pathway through a native FISH-based optical screen.Cell Rep. 2025 Jan 28;44(1):115114. doi: 10.1016/j.celrep.2024.115114. Epub 2024 Dec 26. Cell Rep. 2025. PMID: 39729394 Free PMC article.
-
The TLK-ASF1 histone chaperone pathway plays a critical role in IL-1β-mediated AML progression.Blood. 2024 Jun 27;143(26):2749-2762. doi: 10.1182/blood.2023022079. Blood. 2024. PMID: 38498025 Free PMC article.
References
-
- Min W., Bruhn C., Grigaravicius P., Zhou Z.-W., Li F., Krüger A., Siddeek B., Greulich K.-O., Popp O., Meisezahl C., Calkhoven C. F., Bürkle A., Xu X., Wang Z.-Q., Poly(ADP-ribose) binding to Chk1 at stalled replication forks is required for S-phase checkpoint activation. Nat. Commun. 4, 2993 (2013). - PubMed
-
- Alabert C., Groth A., Chromatin replication and epigenome maintenance. Nat. Rev. Mol. Cell Biol. 13, 153–167 (2012). - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

