Ras-Specific GTPase-Activating Proteins-Structures, Mechanisms, and Interactions
- PMID: 30104198
- PMCID: PMC6396337
- DOI: 10.1101/cshperspect.a031500
Ras-Specific GTPase-Activating Proteins-Structures, Mechanisms, and Interactions
Abstract
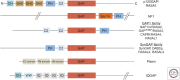
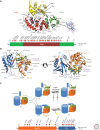
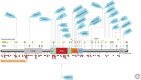
Ras-specific GTPase-activating proteins (RasGAPs) down-regulate the biological activity of Ras proteins by accelerating their intrinsic rate of GTP hydrolysis, basically by a transition state stabilizing mechanism. Oncogenic Ras is commonly not sensitive to RasGAPs caused by interference of mutants with the electronic or steric requirements of the transition state, resulting in up-regulation of activated Ras in respective cells. RasGAPs are modular proteins containing a helical catalytic RasGAP module surrounded by smaller domains that are frequently involved in the subcellular localization or contributing to regulatory features of their host proteins. In this review, we summarize current knowledge about RasGAP structure, mechanism, regulation, and dual-substrate specificity and discuss in some detail neurofibromin, one of the most important negative Ras regulators in cellular growth control and neuronal function.
Copyright © 2019 Cold Spring Harbor Laboratory Press; all rights reserved.
Figures


References
-
- Ahmadian MR, Wiesmuller L, Lautwein A, Bischoff FR, Wittinghofer A. 1996. Structural differences in the minimal catalytic domains of the GTPase-activating proteins p120GAP and neurofibromin. J Biol Chem 271: 16409–16415. - PubMed
-
- Ahmadian MR, Hoffmann U, Goody RS, Wittinghofer A. 1997a. Individual rate constants for the interaction of Ras proteins with GTPase-activating proteins determined by fluorescence spectroscopy. Biochemistry 36: 4535–4541. - PubMed
-
- Ahmadian MR, Mittal R, Hall A, Wittinghofer A. 1997b. Aluminum fluoride associates with the small guanine nucleotide binding proteins. FEBS Lett 408: 315–318. - PubMed
-
- Ahmadian MR, Stege P, Scheffzek K, Wittinghofer A. 1997c. Confirmation of the arginine-finger hypothesis for the GAP-stimulated GTP-hydrolysis reaction of Ras. Nat Struct Biol 4: 686–689. - PubMed
-
- Ahmadian MR, Kiel C, Stege P, Scheffzek K. 2003. Structural fingerprints of the Ras-GTPase activating proteins neurofibromin and p120GAP. J Mol Biol 329: 699–710. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
