A Novel LC System Embeds Analytes in Pre-formed Gradients for Rapid, Ultra-robust Proteomics
- PMID: 30104208
- PMCID: PMC6210218
- DOI: 10.1074/mcp.TIR118.000853
A Novel LC System Embeds Analytes in Pre-formed Gradients for Rapid, Ultra-robust Proteomics
Abstract
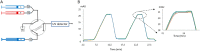
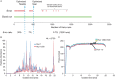
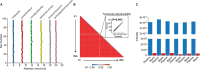
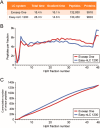
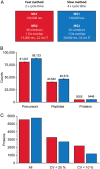
To further integrate mass spectrometry (MS)-based proteomics into biomedical research and especially into clinical settings, high throughput and robustness are essential requirements. They are largely met in high-flow rate chromatographic systems for small molecules but these are not sufficiently sensitive for proteomics applications. Here we describe a new concept that delivers on these requirements while maintaining the sensitivity of current nano-flow LC systems. Low-pressure pumps elute the sample from a disposable trap column, simultaneously forming a chromatographic gradient that is stored in a long storage loop. An auxiliary gradient creates an offset, ensuring the re-focusing of the peptides before the separation on the analytical column by a single high-pressure pump. This simplified design enables robust operation over thousands of sample injections. Furthermore, the steps between injections are performed in parallel, reducing overhead time to a few minutes and allowing analysis of more than 200 samples per day. From fractionated HeLa cell lysates, deep proteomes covering more than 130,000 sequence unique peptides and close to 10,000 proteins were rapidly acquired. Using this data as a library, we demonstrate quantitation of 5200 proteins in only 21 min. Thus, the new system - termed Evosep One - analyzes samples in an extremely robust and high throughput manner, without sacrificing in depth proteomics coverage.
Keywords: Automation; Clinical proteomics; HPLC; High Throughput Screening; Mass Spectrometry; Pre-formed gradient; Robustness; StageTip.
© 2018 Bache et al.
Conflict of interest statement
The authors state that they have potential conflicts of interest regarding this work: NB, OH, LF, OV, are employees of Evosep and MM is an indirect investor in Evosep
Figures




References
-
- Aebersold R., and Mann M. (2016) Mass-spectrometric exploration of proteome structure and function. Nature 537, 347–355 - PubMed
-
- Kulak N. A., Pichler G., Paron I., Nagaraj N., and Mann M. (2014) Minimal, encapsulated proteomic-sample processing applied to copy-number estimation in eukaryotic cells. Nat. Methods 11, 319–324 - PubMed
-
- Cox J., and Mann M. (2008) MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nat. Biotechnol. 26, 1367–1372 - PubMed
-
- Bekker-Jensen D.B., Kelstrup C. D., Batth T. S., Larsen S. C., Haldrup C., Bramsen J. B., Sørensen K. D., Høyer S., Ørntoft T. F., Andersen C. L., Nielsen M. L., and Olsen J. V. (2017) An optimized shotgun strategy for the rapid generation of comprehensive human proteomes. Cell Systems 4, 587–599 - PMC - PubMed
-
- Kelstrup C. D., Bekker-Jensen D. B., Arrey T. N., Hogrebe A., Harder A., and Olsen J. V. (2018) Performance evaluation of the Q Exactive HF-X for shotgun proteomics. J Proteome Res, 17, 727–738 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

