Highly efficient cellular uptake of a cell-penetrating peptide (CPP) derived from the capsid protein of porcine circovirus type 2
- PMID: 30108178
- PMCID: PMC6166719
- DOI: 10.1074/jbc.RA118.004823
Highly efficient cellular uptake of a cell-penetrating peptide (CPP) derived from the capsid protein of porcine circovirus type 2
Abstract
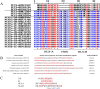
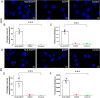
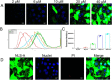
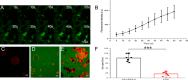
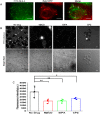
Porcine circovirus type 2 (PCV2) is one of the smallest, nonenveloped, single-stranded DNA viruses. The PCV2 capsid protein (Cap) is the sole viral structural protein and main antigenic determinant. Previous sequence analysis has revealed that the N terminus of the PCV2 Cap contains a nuclear localization signal (NLS) enriched in positively charged residues. Here, we report that PCV2's NLS can function as a cell-penetrating peptide (CPP). We observed that this NLS can carry macromolecules, e.g. enhanced GFP (EGFP), into cells when they are fused to the NLS, indicating that it can function as a CPP, similar to the classical CPP derived from HIV type 1 transactivator of transcription protein (HIV TAT). We also found that the first 17 residues of the NLS (NLS-A) have a key role in cellular uptake. In addition to entering cells via multiple endocytic processes, NLS-A was also rapidly internalized via direct translocation enabled by increased membrane permeability and was evenly distributed throughout cells when its concentration in cell cultures was ≥10 μm Of note, cellular NLS-A uptake was ∼10 times more efficient than that of HIV TAT. We inferred that the externalized NLS of the PCV2 Cap may accumulate to a high concentration (≥10 μm) at a local membrane area, increasing membrane permeability to facilitate viral entry into the cell to release its genome into a viral DNA reproduction center. We conclude that NLS-A has potential as a versatile vehicle for shuttling foreign molecules into cells, including pharmaceuticals for therapeutic interventions.
Keywords: Circoviridae; capsid protein; cell-penetrating peptide (CPP); cellular uptake; endocytosis; intracellular trafficking; membrane; membrane permeability; nuclear localization signal (NLS); permeability; porcine circovirus virus (PCV); protein delivery; transport vector.
© 2018 Yu et al.
Conflict of interest statement
The authors declare that they have no conflicts of interest with the contents of this article
Figures

Similar articles
-
The Carboxyl Terminus of the Porcine Circovirus Type 2 Capsid Protein Is Critical to Virus-Like Particle Assembly, Cell Entry, and Propagation.J Virol. 2020 Apr 16;94(9):e00042-20. doi: 10.1128/JVI.00042-20. Print 2020 Apr 16. J Virol. 2020. PMID: 32075927 Free PMC article.
-
Nuclear localization signal regulates porcine circovirus type 2 capsid protein nuclear export through phosphorylation.Virus Res. 2018 Feb 15;246:12-22. doi: 10.1016/j.virusres.2017.12.012. Epub 2017 Dec 30. Virus Res. 2018. PMID: 29292065
-
Functional exchangeability of the nuclear localization signal (NLS) of capsid protein between PCV1 and PCV2 in vitro: Implications for the role of NLS in viral replication.Virol J. 2011 Jul 6;8:341. doi: 10.1186/1743-422X-8-341. Virol J. 2011. PMID: 21733152 Free PMC article.
-
Structural Perspectives of Beak and Feather Disease Virus and Porcine Circovirus Proteins.Viral Immunol. 2021 Jan-Feb;34(1):49-59. doi: 10.1089/vim.2020.0097. Epub 2020 Dec 4. Viral Immunol. 2021. PMID: 33275868 Review.
-
Porcine Circovirus Type 2 (PCV2) Vaccines in the Context of Current Molecular Epidemiology.Viruses. 2017 May 6;9(5):99. doi: 10.3390/v9050099. Viruses. 2017. PMID: 28481275 Free PMC article. Review.
Cited by
-
Novel Lentivirus-Based Method for Rapid Selection of Inhibitory Nanobody against PRRSV.Viruses. 2020 Feb 19;12(2):229. doi: 10.3390/v12020229. Viruses. 2020. PMID: 32092857 Free PMC article.
-
Epidemiological investigation and analysis of the infection of porcine circovirus in Xinjiang.Virol J. 2024 Sep 27;21(1):230. doi: 10.1186/s12985-024-02504-w. Virol J. 2024. PMID: 39334389 Free PMC article.
-
Bound Protein- and Peptide-Based Strategies for Adeno-Associated Virus Vector-Mediated Gene Therapy: Where Do We Stand Now?Hum Gene Ther. 2020 Nov;31(21-22):1146-1154. doi: 10.1089/hum.2020.193. Epub 2020 Oct 22. Hum Gene Ther. 2020. PMID: 32940063 Free PMC article.
-
Mitochondrial Localization Signal of Porcine Circovirus Type 2 Capsid Protein Plays a Critical Role in Cap-Induced Apoptosis.Vet Sci. 2021 Nov 10;8(11):272. doi: 10.3390/vetsci8110272. Vet Sci. 2021. PMID: 34822645 Free PMC article.
-
MeCP2 is a naturally supercharged protein with cell membrane transduction capabilities.Protein Sci. 2024 Oct;33(10):e5170. doi: 10.1002/pro.5170. Protein Sci. 2024. PMID: 39276009 Free PMC article.
References
-
- Tischer I., Rasch R., and Tochtermann G. (1974) Characterization of papovavirus-and picornavirus-like particles in permanent pig kidney cell lines. Zentralbl. Bakteriol. Orig. A 226, 153–167 - PubMed
-
- Palinski R., Piñeyro P., Shang P., Yuan F., Guo R., Fang Y., Byers E., and Hause B. M. (2017) A novel porcine circovirus distantly related to known circoviruses is associated with porcine dermatitis and nephropathy syndrome and reproductive failure. J. Virol. 91, e01879–16 10.1128/JVI.01879-16 - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

