Sequence Variation of Epstein-Barr Virus: Viral Types, Geography, Codon Usage, and Diseases
- PMID: 30111570
- PMCID: PMC6206488
- DOI: 10.1128/JVI.01132-18
Sequence Variation of Epstein-Barr Virus: Viral Types, Geography, Codon Usage, and Diseases
Abstract
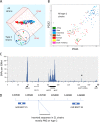
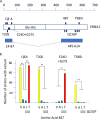
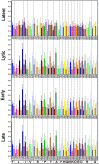
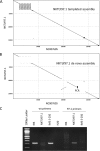
One hundred thirty-eight new Epstein-Barr virus (EBV) genome sequences have been determined. One hundred twenty-five of these and 116 from previous reports were combined to produce a multiple-sequence alignment of 241 EBV genomes, which we have used to analyze variation within the viral genome. The type 1/type 2 classification of EBV remains the major form of variation and is defined mostly by EBNA2 and EBNA3, but the type 2 single-nucleotide polymorphisms (SNPs) at the EBNA3 locus extend into the adjacent gp350 and gp42 genes, whose products mediate infection of B cells by EBV. A small insertion within the BART microRNA region of the genome was present in 21 EBV strains. EBV from saliva of U.S. patients with chronic active EBV infection aligned with the wild-type EBV genome with no evidence of WZhet rearrangements. The V3 polymorphism in the Zp promoter for BZLF1 was found to be frequent in nasopharyngeal carcinoma cases from both Hong Kong and Indonesia. Codon usage was found to differ between latent and lytic cycle EBV genes, and the main forms of variation of the EBNA1 protein have been identified.IMPORTANCE Epstein-Barr virus causes most cases of infectious mononucleosis and posttransplant lymphoproliferative disease. It contributes to several types of cancer, including Hodgkin's lymphoma, Burkitt's lymphoma, diffuse large B cell lymphoma, nasopharyngeal carcinoma, and gastric carcinoma. EBV genome variation is important because some of the diseases associated with EBV have very different incidences in different populations and geographic regions, and differences in the EBV genome might contribute to these diseases. Some specific EBV genome alterations that appear to be significant in EBV-associated cancers are already known, and current efforts to make an EBV vaccine and antiviral drugs should also take account of sequence differences in the proteins used as targets.
Keywords: Epstein-Barr virus.
Copyright © 2018 Correia et al.
Figures


References
-
- Kelly GL, Long HM, Stylianou J, Thomas WA, Leese A, Bell AI, Bornkamm GW, Mautner J, Rickinson AB, Rowe M. 2009. An Epstein-Barr virus anti-apoptotic protein constitutively expressed in transformed cells and implicated in Burkitt lymphomagenesis: the Wp/BHRF1 link. PLoS Pathog 5:e1000341. doi:10.1371/journal.ppat.1000341. - DOI - PMC - PubMed
-
- White RE, Ramer PC, Naresh KN, Meixlsperger S, Pinaud L, Rooney C, Savoldo B, Coutinho R, Bodor C, Gribben J, Ibrahim HA, Bower M, Nourse JP, Gandhi MK, Middeldorp J, Cader FZ, Murray P, Munz C, Allday MJ. 2012. EBNA3B-deficient EBV promotes B cell lymphomagenesis in humanized mice and is found in human tumors. J Clin Investig 122:1487–1502. doi:10.1172/JCI58092. - DOI - PMC - PubMed
-
- Palser AL, Grayson NE, White RE, Corton C, Correia S, Ba Abdullah MM, Watson SJ, Cotten M, Arrand JR, Murray PG, Allday MJ, Rickinson AB, Young LS, Farrell PJ, Kellam P. 2015. Genome diversity of Epstein-Barr virus from multiple tumor types and normal infection. J Virol 89:5222–5237. doi:10.1128/JVI.03614-14. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

