The miR-96 and RARγ signaling axis governs androgen signaling and prostate cancer progression
- PMID: 30120411
- PMCID: PMC6336686
- DOI: 10.1038/s41388-018-0450-6
The miR-96 and RARγ signaling axis governs androgen signaling and prostate cancer progression
Abstract
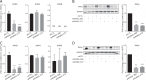
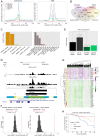
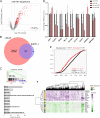
Expression levels of retinoic acid receptor gamma (NR1B3/RARG, encodes RARγ) are commonly reduced in prostate cancer (PCa). Therefore, we sought to establish the cellular and gene regulatory consequences of reduced RARγ expression, and determine RARγ regulatory mechanisms. RARG shRNA approaches in non-malignant (RWPE-1 and HPr1-AR) and malignant (LNCaP) prostate models revealed that reducing RARγ levels, rather than adding exogenous retinoid ligand, had the greatest impact on prostate cell viability and gene expression. ChIP-Seq defined the RARγ cistrome, which was significantly enriched at active enhancers associated with AR binding sites. Reflecting a significant genomic role for RARγ to regulate androgen signaling, RARγ knockdown in HPr1-AR cells significantly regulated the magnitude of the AR transcriptome. RARγ downregulation was explained by increased miR-96 in PCa cell and mouse models, and TCGA PCa cohorts. Biochemical approaches confirmed that miR-96 directly regulated RARγ expression and function. Capture of the miR-96 targetome by biotin-miR-96 identified that RARγ and a number of RARγ interacting co-factors including TACC1 were all targeted by miR-96, and expression of these genes were prominently altered, positively and negatively, in the TCGA-PRAD cohort. Differential gene expression analyses between tumors in the TCGA-PRAD cohort with lower quartile expression levels of RARG and TACC1 and upper quartile miR-96, compared to the reverse, identified a gene network including several RARγ target genes (e.g., SOX15) that significantly associated with worse disease-free survival (hazard ratio 2.23, 95% CI 1.58 to 2.88, p = 0.015). In summary, miR-96 targets a RARγ network to govern AR signaling, PCa progression and disease outcome.
Conflict of interest statement
The authors declare that they have no conflict of interest.
Figures





Comment in
-
miR-96 influences RARγ expression.Nat Rev Urol. 2018 Nov;15(11):656-657. doi: 10.1038/s41585-018-0088-8. Nat Rev Urol. 2018. PMID: 30194359 No abstract available.
Similar articles
-
Identification of novel androgen receptor target genes in prostate cancer.Mol Cancer. 2007 Jun 6;6:39. doi: 10.1186/1476-4598-6-39. Mol Cancer. 2007. PMID: 17553165 Free PMC article.
-
Crosstalk between epithelial-mesenchymal transition and castration resistance mediated by Twist1/AR signaling in prostate cancer.Endocr Relat Cancer. 2015 Dec;22(6):889-900. doi: 10.1530/ERC-15-0225. Epub 2015 Aug 26. Endocr Relat Cancer. 2015. PMID: 26311513
-
Androgen-regulated miR-32 targets BTG2 and is overexpressed in castration-resistant prostate cancer.Oncogene. 2012 Oct 11;31(41):4460-71. doi: 10.1038/onc.2011.624. Epub 2012 Jan 23. Oncogene. 2012. PMID: 22266859
-
Androgen receptor and prostate cancer stem cells: biological mechanisms and clinical implications.Endocr Relat Cancer. 2015 Dec;22(6):T209-20. doi: 10.1530/ERC-15-0217. Epub 2015 Aug 18. Endocr Relat Cancer. 2015. PMID: 26285606 Free PMC article. Review.
-
Androgen receptor action in hormone-dependent and recurrent prostate cancer.J Cell Biochem. 2006 Oct 1;99(2):362-72. doi: 10.1002/jcb.20811. J Cell Biochem. 2006. PMID: 16619264 Review.
Cited by
-
MicroRNA Dysregulation in Prostate Cancer.Pharmgenomics Pers Med. 2022 Mar 10;15:177-193. doi: 10.2147/PGPM.S348565. eCollection 2022. Pharmgenomics Pers Med. 2022. PMID: 35300057 Free PMC article.
-
Interaction between Non-Coding RNAs and Androgen Receptor with an Especial Focus on Prostate Cancer.Cells. 2021 Nov 16;10(11):3198. doi: 10.3390/cells10113198. Cells. 2021. PMID: 34831421 Free PMC article. Review.
-
African American Prostate Cancer Displays Quantitatively Distinct Vitamin D Receptor Cistrome-transcriptome Relationships Regulated by BAZ1A.Cancer Res Commun. 2023 Apr 18;3(4):621-639. doi: 10.1158/2767-9764.CRC-22-0389. eCollection 2023 Apr. Cancer Res Commun. 2023. PMID: 37082578 Free PMC article.
-
miR-769-5p is associated with prostate cancer recurrence and modulates proliferation and apoptosis of cancer cells.Exp Ther Med. 2021 Apr;21(4):335. doi: 10.3892/etm.2021.9766. Epub 2021 Feb 8. Exp Ther Med. 2021. PMID: 33732308 Free PMC article.
-
Dynamic patterns of DNA methylation in the normal prostate epithelial differentiation program are targets of aberrant methylation in prostate cancer.Sci Rep. 2021 Jun 1;11(1):11405. doi: 10.1038/s41598-021-91037-1. Sci Rep. 2021. PMID: 34075163 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials

