Cysteine residues in mitochondrial intermembrane space proteins: more than just import
- PMID: 30129023
- PMCID: PMC6346212
- DOI: 10.1111/bph.14480
Cysteine residues in mitochondrial intermembrane space proteins: more than just import
Abstract
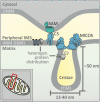
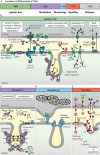
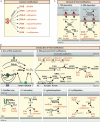
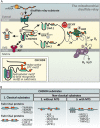
The intermembrane space (IMS) is a very small mitochondrial sub-compartment with critical relevance for many cellular processes. IMS proteins fulfil important functions in transport of proteins, lipids, metabolites and metal ions, in signalling, in metabolism and in defining the mitochondrial ultrastructure. Our understanding of the IMS proteome has become increasingly refined although we still lack information on the identity and function of many of its proteins. One characteristic of many IMS proteins are conserved cysteines. Different post-translational modifications of these cysteine residues can have critical roles in protein function, localization and/or stability. The close localization to different ROS-producing enzyme systems, a dedicated machinery for oxidative protein folding, and a unique equipment with antioxidative systems, render the careful balancing of the redox and modification states of the cysteine residues, a major challenge in the IMS. In this review, we discuss different functions of human IMS proteins, the involvement of cysteine residues in these functions, the consequences of cysteine modifications and the consequences of cysteine mutations or defects in the machinery for disulfide bond formation in terms of human health. LINKED ARTICLES: This article is part of a themed section on Chemical Biology of Reactive Sulfur Species. To view the other articles in this section visit http://onlinelibrary.wiley.com/doi/10.1111/bph.v176.4/issuetoc.
© 2018 The British Pharmacological Society.
Figures

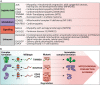
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

