Modeling human somite development and fibrodysplasia ossificans progressiva with induced pluripotent stem cells
- PMID: 30139810
- PMCID: PMC6124548
- DOI: 10.1242/dev.165431
Modeling human somite development and fibrodysplasia ossificans progressiva with induced pluripotent stem cells
Abstract
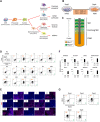
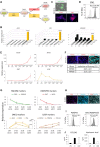
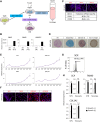
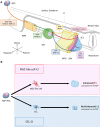
Somites (SMs) comprise a transient stem cell population that gives rise to multiple cell types, including dermatome (D), myotome (MYO), sclerotome (SCL) and syndetome (SYN) cells. Although several groups have reported induction protocols for MYO and SCL from pluripotent stem cells, no studies have demonstrated the induction of SYN and D from SMs. Here, we report systematic induction of these cells from human induced pluripotent stem cells (iPSCs) under chemically defined conditions. We also successfully induced cells with differentiation capacities similar to those of multipotent mesenchymal stromal cells (MSC-like cells) from SMs. To evaluate the usefulness of these protocols, we conducted disease modeling of fibrodysplasia ossificans progressiva (FOP), an inherited disease that is characterized by heterotopic endochondral ossification in soft tissues after birth. Importantly, FOP-iPSC-derived MSC-like cells showed enhanced chondrogenesis, whereas FOP-iPSC-derived SCL did not, possibly recapitulating normal embryonic skeletogenesis in FOP and cell-type specificity of FOP phenotypes. These results demonstrate the usefulness of multipotent SMs for disease modeling and future cell-based therapies.
Keywords: Differentiation; Disease modeling; Fibrodysplasia ossificans progressiva; Induced pluripotent stem cells; Paraxial mesoderm.
© 2018. Published by The Company of Biologists Ltd.
Conflict of interest statement
Competing interestsThe authors declare no competing or financial interests.
Figures


Similar articles
-
An mTOR Signaling Modulator Suppressed Heterotopic Ossification of Fibrodysplasia Ossificans Progressiva.Stem Cell Reports. 2018 Nov 13;11(5):1106-1119. doi: 10.1016/j.stemcr.2018.10.007. Epub 2018 Nov 1. Stem Cell Reports. 2018. PMID: 30392977 Free PMC article.
-
Activin-A enhances mTOR signaling to promote aberrant chondrogenesis in fibrodysplasia ossificans progressiva.J Clin Invest. 2017 Sep 1;127(9):3339-3352. doi: 10.1172/JCI93521. Epub 2017 Jul 31. J Clin Invest. 2017. PMID: 28758906 Free PMC article.
-
Neofunction of ACVR1 in fibrodysplasia ossificans progressiva.Proc Natl Acad Sci U S A. 2015 Dec 15;112(50):15438-43. doi: 10.1073/pnas.1510540112. Epub 2015 Nov 30. Proc Natl Acad Sci U S A. 2015. PMID: 26621707 Free PMC article.
-
Application of human induced pluripotent stem cells to model fibrodysplasia ossificans progressiva.Bone. 2018 Apr;109:162-167. doi: 10.1016/j.bone.2017.07.003. Epub 2017 Jul 14. Bone. 2018. PMID: 28716551 Free PMC article. Review.
-
Using Human Induced Pluripotent Stem Cells to Model Skeletal Diseases.Methods Mol Biol. 2016;1353:101-18. doi: 10.1007/7651_2014_171. Methods Mol Biol. 2016. PMID: 25549831 Review.
Cited by
-
Single Cell Transcriptomics-Informed Induced Pluripotent Stem Cells Differentiation to Tenogenic Lineage.bioRxiv [Preprint]. 2024 Jul 10:2023.04.10.536240. doi: 10.1101/2023.04.10.536240. bioRxiv. 2024. PMID: 37090543 Free PMC article. Preprint.
-
Epigenetic mechanisms in stem cell therapies for achilles tendinopathy.Front Cell Dev Biol. 2025 Mar 13;13:1516250. doi: 10.3389/fcell.2025.1516250. eCollection 2025. Front Cell Dev Biol. 2025. PMID: 40181824 Free PMC article. Review.
-
Interspecies control of development during mammalian gastrulation.Emerg Top Life Sci. 2023 Dec 18;7(4):397-408. doi: 10.1042/ETLS20230083. Emerg Top Life Sci. 2023. PMID: 37933589 Free PMC article. Review.
-
Application of mesenchymal stem/stromal cells in periodontal regeneration: Opportunities and challenges.Jpn Dent Sci Rev. 2024 Dec;60:95-108. doi: 10.1016/j.jdsr.2024.01.001. Epub 2024 Jan 25. Jpn Dent Sci Rev. 2024. PMID: 38314143 Free PMC article. Review.
-
Mesodermal Derivatives of Pluripotent Stem Cells Route to Scarless Healing.Int J Mol Sci. 2023 Jul 26;24(15):11945. doi: 10.3390/ijms241511945. Int J Mol Sci. 2023. PMID: 37569321 Free PMC article. Review.
References
-
- Bernardo A. S., Faial T., Gardner L., Niakan K. K., Ortmann D., Senner C. E., Callery E. M., Trotter M. W., Hemberger M., Smith J. C. et al. (2011). BRACHYURY and CDX2 mediate BMP-induced differentiation of human and mouse pluripotent stem cells into embryonic and extraembryonic lineages. Cell Stem Cell 9, 144-155. 10.1016/j.stem.2011.06.015 - DOI - PMC - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

