Host functions used by hepatitis B virus to complete its life cycle: Implications for developing host-targeting agents to treat chronic hepatitis B
- PMID: 30145242
- PMCID: PMC6193490
- DOI: 10.1016/j.antiviral.2018.08.014
Host functions used by hepatitis B virus to complete its life cycle: Implications for developing host-targeting agents to treat chronic hepatitis B
Abstract
Similar to other mammalian viruses, the life cycle of hepatitis B virus (HBV) is heavily dependent upon and regulated by cellular (host) functions. These cellular functions can be generally placed in to two categories: (a) intrinsic host restriction factors and innate defenses, which must be evaded or repressed by the virus; and (b) gene products that provide functions necessary for the virus to complete its life cycle. Some of these functions may apply to all viruses, but some may be specific to HBV. In certain cases, the virus may depend upon the host function much more than does the host itself. Knowing which host functions regulate the different steps of a virus' life cycle, can lead to new antiviral targets and help in developing novel treatment strategies, in addition to improving a fundamental understanding of viral pathogenesis. Therefore, in this review we will discuss known host factors which influence key steps of HBV life cycle, and further elucidate therapeutic interventions targeting host-HBV interactions.
Keywords: Antiviral agents; Direct acting antiviral agents; Hepatitis; Hepatitis B virus (HBV); Host targeting agents; Liver; Virus-host interaction.
Copyright © 2018. Published by Elsevier B.V.
Figures



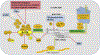
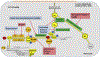

References
-
- Alter H, Block T, Brown N, Brownstein A, Brosgart C, Chang KM, Chen PJ, Chisari FV, Cohen C, El-Serag H, Feld J, Gish R, Glenn J, Greten T, Guo H, Guo JT, Hoshida Y, Hu J, Kowdley KV, Li W, Liang J, Locarnini S, Lok AS, Mason W, McMahon B, Mehta A, Perrillo R, Revill P, Rice CM, Rinaudo J, Schinazi R, Seeger C, Shetty K, Tavis J, Zoulim F, 2018. A research agenda for curing chronic hepatitis B virus infection. Hepatology 67, 1127–1131. - PMC - PubMed
-
- Bayliss J, Lim L, Thompson AJ, Desmond P, Angus P, Locarnini S, Revill PA, 2013. Hepatitis B virus splicing is enhanced prior to development of hepatocellular carcinoma. J. Hepatol 59, 1022–1028. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases

