Molecular mechanism of cardiolipin-mediated assembly of respiratory chain supercomplexes
- PMID: 30155091
- PMCID: PMC6014297
- DOI: 10.1039/c5sc04664e
Molecular mechanism of cardiolipin-mediated assembly of respiratory chain supercomplexes
Abstract
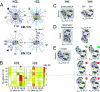
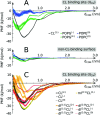
Mitochondria produce most of the ATP consumed by cells through the respiratory chain in their inner membrane. This process involves protein complexes assembled into larger structures, the respiratory supercomplexes (SCs). Cardiolipin (CL), the mitochondrial signature phospholipid, is crucial for the structural and functional integrity of these SCs, but it is as yet unclear by what mechanism it operates. Our data disclose the mechanism for bulk CL in gluing SCs, steering their formation, and suggest how it may stabilize specific interfaces. We describe self-assembly molecular dynamics simulations of 9 cytochrome bc1 (CIII) dimers and 27 cytochrome c oxidase (CIV) monomers from bovine heart mitochondria embedded in a CL-containing model lipid bilayer, aimed at mimicking the crowdedness and complexity of mitochondrial membranes. The simulations reveal a large diversity of interfaces, including those of existing experimental CIII/CIV SC models and an alternative interface with CIV rotated by 180°. SC interfaces enclose 4 to 12 CLs, a ∼10 fold enrichment from the bulk. Half of these CLs glue complexes together using CL binding sites at the surface of both complexes. Free energy calculations demonstrate a larger CL binding strength, compared to other mitochondrial lipids, that is exclusive to these binding sites and results from non-additive electrostatic and van der Waals forces. This study provides a key example of the ability of lipids to selectively mediate protein-protein interactions by altering all ranges of forces, lubricate protein interfaces and act as traffic control agents steering proteins together.
Figures




References
LinkOut - more resources
Full Text Sources
Other Literature Sources

