Modeling Diversity: Do Homogeneous Laboratory Strains Limit Discovery?
- PMID: 30166218
- PMCID: PMC6610874
- DOI: 10.1016/j.tim.2018.08.002
Modeling Diversity: Do Homogeneous Laboratory Strains Limit Discovery?
Abstract
The outcome of chronic infections is highly variable. The heterogeneous disease outcomes in natural populations differ from genetically homogeneous infection models. Here, we use tuberculosis as a 'case study' to contrast the genetic landscape in natural populations with standard infection models, discussing new strategies to bridge this gap.
Keywords: Collaborative Cross; genetic diversity; host genetics; infectious diseases; mouse models; tuberculosis.
Copyright © 2018 Elsevier Ltd. All rights reserved.
Figures
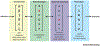
References
-
- Mazurek GH et al. Updated guidelines for using Interferon Gamma Release Assays to detect Mycobacterium tuberculosis infection - United States, 2010., MMWR. Recommendations and reports: Morbidity and mortality weekly report. Recommendations and reports, 59 25-June-(2010), 1–25 - PubMed
-
- Hernández-Pando R et al. (2012) Use of mouse models to study the variability in virulence associated with specific genotypic lineages of Mycobacterium tuberculosis. Infect. Genet. Evol 12, 725–731 - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Miscellaneous

