Multitrait genome association analysis identifies new susceptibility genes for human anthropometric variation in the GCAT cohort
- PMID: 30166351
- PMCID: PMC6252362
- DOI: 10.1136/jmedgenet-2018-105437
Multitrait genome association analysis identifies new susceptibility genes for human anthropometric variation in the GCAT cohort
Abstract
Background: Heritability estimates have revealed an important contribution of SNP variants for most common traits; however, SNP analysis by single-trait genome-wide association studies (GWAS) has failed to uncover their impact. In this study, we applied a multitrait GWAS approach to discover additional factor of the missing heritability of human anthropometric variation.
Methods: We analysed 205 traits, including diseases identified at baseline in the GCAT cohort (Genomes For Life- Cohort study of the Genomes of Catalonia) (n=4988), a Mediterranean adult population-based cohort study from the south of Europe. We estimated SNP heritability contribution and single-trait GWAS for all traits from 15 million SNP variants. Then, we applied a multitrait-related approach to study genome-wide association to anthropometric measures in a two-stage meta-analysis with the UK Biobank cohort (n=336 107).
Results: Heritability estimates (eg, skin colour, alcohol consumption, smoking habit, body mass index, educational level or height) revealed an important contribution of SNP variants, ranging from 18% to 77%. Single-trait analysis identified 1785 SNPs with genome-wide significance threshold. From these, several previously reported single-trait hits were confirmed in our sample with LINC01432 (p=1.9×10-9) variants associated with male baldness, LDLR variants with hyperlipidaemia (ICD-9:272) (p=9.4×10-10) and variants in IRF4 (p=2.8×10-57), SLC45A2 (p=2.2×10-130), HERC2 (p=2.8×10-176), OCA2 (p=2.4×10-121) and MC1R (p=7.7×10-22) associated with hair, eye and skin colour, freckling, tanning capacity and sun burning sensitivity and the Fitzpatrick phototype score, all highly correlated cross-phenotypes. Multitrait meta-analysis of anthropometric variation validated 27 loci in a two-stage meta-analysis with a large British ancestry cohort, six of which are newly reported here (p value threshold <5×10-9) at ZRANB2-AS2, PIK3R1, EPHA7, MAD1L1, CACUL1 and MAP3K9.
Conclusion: Considering multiple-related genetic phenotypes improve associated genome signal detection. These results indicate the potential value of data-driven multivariate phenotyping for genetic studies in large population-based cohorts to contribute to knowledge of complex traits.
Keywords: cohort; complex traits; gwas; multitrait; phenome.
© Author(s) (or their employer(s)) 2018. Re-use permitted under CC BY-NC. No commercial re-use. See rights and permissions. Published by BMJ.
Conflict of interest statement
Competing interests: None declared.
Figures
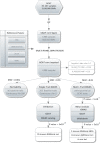

References
-
- Eurostat Statistics Explained. Mortality and life expectancy statistics, 2016. http://ec.europa.eu/eurostat/statistics-explained/index.php/Mortality_an...
-
- Riboli E, Hunt KJ, Slimani N, Ferrari P, Norat T, Fahey M, Charrondière UR, Hémon B, Casagrande C, Vignat J, Overvad K, Tjønneland A, Clavel-Chapelon F, Thiébaut A, Wahrendorf J, Boeing H, Trichopoulos D, Trichopoulou A, Vineis P, Palli D, Bueno-De-Mesquita HB, Peeters PH, Lund E, Engeset D, González CA, Barricarte A, Berglund G, Hallmans G, Day NE, Key TJ, Kaaks R, Saracci R. European Prospective Investigation into Cancer and Nutrition (EPIC): study populations and data collection. Public Health Nutr 2002;5:1113–24. 10.1079/PHN2002394 - DOI - PubMed
-
- Manolio TA, Collins FS, Cox NJ, Goldstein DB, Hindorff LA, Hunter DJ, McCarthy MI, Ramos EM, Cardon LR, Chakravarti A, Cho JH, Guttmacher AE, Kong A, Kruglyak L, Mardis E, Rotimi CN, Slatkin M, Valle D, Whittemore AS, Boehnke M, Clark AG, Eichler EE, Gibson G, Haines JL, Mackay TF, McCarroll SA, Visscher PM. Finding the missing heritability of complex diseases. Nature 2009;461:747–53. 10.1038/nature08494 - DOI - PMC - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous
