Terminal hairpin in oligonucleotide dominantly prioritizes intramolecular cyclization by T4 ligase over intermolecular polymerization: an exclusive methodology for producing ssDNA rings
- PMID: 30169701
- PMCID: PMC6294566
- DOI: 10.1093/nar/gky769
Terminal hairpin in oligonucleotide dominantly prioritizes intramolecular cyclization by T4 ligase over intermolecular polymerization: an exclusive methodology for producing ssDNA rings
Abstract
When oligonucleotide bearing a hairpin near either its 3'- or 5'-end was treated with T4 DNA ligase, the intramolecular cyclization dominantly proceeded and its monomeric cyclic ring was obtained in extremely high selectivity. The selectivity was hardly dependent on the concentration of the oligonucleotide, and thus it could be added in one portion to the mixture at the beginning of the reaction. Without the hairpin, however, the formation of polymeric byproducts was dominant under the same conditions. Hairpin-bearing oligonucleotides primarily take the folded form, and the enzymatically reactive species (its open form) is minimal. As the result, the intermolecular reactions are efficiently suppressed due to both thermodynamic and kinetic factors. The 'terminal hairpin strategy' was applicable to large-scale preparation of a variety of DNA rings. The combination of this methodology with 'diluted buffer strategy', developed previously, is still more effective for the purpose. When large amount of l-DNA bearing a terminal hairpin (e.g. 40 μM) was treated in a diluted ligase buffer (0.1× buffer) with T4 DNA ligase, the DNA ring was prepared in 100% selectivity. Even at [l-DNA]0 = 100 μM in 0.1× buffer, the DNA ring was also obtained in pure form, simply by removing tiny quantity of linear byproducts by Exonuclease I.
Figures
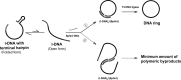
 ).
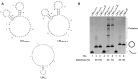
).
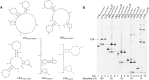


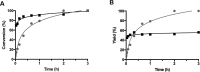

References
-
- Jonathan B., Turberfield A.J.. DNA nanomachines. Nat. Nanotechnol. 2007; 2:275–284. - PubMed
-
- Teller C., Willner I.. Functional nucleic acid nanostructures and DNA machines. Curr. Opin. Biotechnol. 2010; 21:376–391. - PubMed
-
- Komiyama M., Yoshimoto K., Sisido M., Ariga K.. Chemistry can make strict and fuzzy controls for bio-systems: DNA nanoarchitectonics and cell-macromolecular nanoarchitectonics. Bull. Chem. Soc. Jpn. 2017; 90:967–1004.
-
- He Y., Ye T., Su M., Zhang C., Ribbe A.E., Jiang W., Mao C.. Hierarchical self-assembly of DNA into symmetric supramolecular polyhedra. Nature. 2008; 452:198–201. - PubMed
-
- Beissenhirtz M.K., Itamar W.. DNA-based machines. Org. Biomol. Chem. 2006; 4:3392–3401. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

