Visualizing Genetic Variants, Short Targets, and Point Mutations in the Morphological Tissue Context with an RNA In Situ Hybridization Assay
- PMID: 30176002
- PMCID: PMC6126797
- DOI: 10.3791/58097
Visualizing Genetic Variants, Short Targets, and Point Mutations in the Morphological Tissue Context with an RNA In Situ Hybridization Assay
Abstract
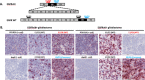
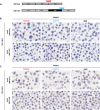
Because precision medicine is highly dependent on the accurate detection of biomarkers, there is an increasing need for standardized and robust technologies that measure RNA biomarkers in situ in clinical specimens. While grind-and-bind assays like RNAseq and quantitative RT-PCR enable highly sensitive gene expression measurements, they also require RNA extraction and thus prevent valuable expression analysis within the morphological tissue context. The in situ hybridization (ISH) assay described here can detect RNA target sequences as short as 50 nucleotides at single-nucleotide resolution and at the single-cell level. This assay is complementary to the previously developed commercial assay and enables sensitive and specific in situ detection of splice variants, short targets, and point mutations within the tissue. In this protocol, probes were designed to target unique exon junctions for two clinically important splice variants, EGFRvIII and METΔ14. The detection of short target sequences was demonstrated by the specific detection of CDR3 sequences of T-cell receptors α and β in the Jurkat T-cell line. Also shown is the utility of this ISH assay for the distinction of RNA target sequences at single-nucleotide resolution (point mutations) through the visualization of EGFR L858R and KRAS G12A single-nucleotide variations in cell lines using automated staining platforms. In summary, the protocol shows a specialized RNA ISH assay that enables the detection of splice variants, short sequences, and mutations in situ for manual performance and on automated stainers.
Similar articles
-
A Novel Ultrasensitive In Situ Hybridization Approach to Detect Short Sequences and Splice Variants with Cellular Resolution.Mol Neurobiol. 2018 Jul;55(7):6169-6181. doi: 10.1007/s12035-017-0834-6. Epub 2017 Dec 20. Mol Neurobiol. 2018. PMID: 29264769 Free PMC article.
-
Quantitative ultrasensitive bright-field RNA in situ hybridization with RNAscope.Methods Mol Biol. 2014;1211:201-12. doi: 10.1007/978-1-4939-1459-3_16. Methods Mol Biol. 2014. PMID: 25218387
-
Branched Chain RNA In Situ Hybridization for Androgen Receptor Splice Variant AR-V7 as a Prognostic Biomarker for Metastatic Castration-Sensitive Prostate Cancer.Clin Cancer Res. 2017 Jan 15;23(2):363-369. doi: 10.1158/1078-0432.CCR-16-0237. Epub 2016 Jul 20. Clin Cancer Res. 2017. PMID: 27440270
-
[RNA in situ hybridization: technology, potential, and fields of application].Pathologe. 2020 Nov;41(6):563-573. doi: 10.1007/s00292-020-00839-z. Pathologe. 2020. PMID: 32997158 Review. German.
-
In situ polymerase chain reaction: general methodology and recent advances.Verh Dtsch Ges Pathol. 1994;78:146-52. Verh Dtsch Ges Pathol. 1994. PMID: 7533976 Review.
Cited by
-
Mutant p53 induces Golgi tubulo-vesiculation driving a prometastatic secretome.Nat Commun. 2020 Aug 7;11(1):3945. doi: 10.1038/s41467-020-17596-5. Nat Commun. 2020. PMID: 32770028 Free PMC article.
-
Detection and Quantification of Multiple RNA Sequences Using Emerging Ultrasensitive Fluorescent In Situ Hybridization Techniques.Curr Protoc Neurosci. 2019 Apr;87(1):e63. doi: 10.1002/cpns.63. Epub 2019 Feb 21. Curr Protoc Neurosci. 2019. PMID: 30791216 Free PMC article.
-
Cost-Efficient and Easy to Perform PCR-Based Assay to Identify Met Exon 14 Skipping in Formalin-Fixed Paraffin-Embedded (FFPE) Non-Small Cell Lung Cancer (NSCLC) Samples.Diagnostics (Basel). 2019 Jan 18;9(1):13. doi: 10.3390/diagnostics9010013. Diagnostics (Basel). 2019. PMID: 30669306 Free PMC article.
-
EGFR T790M detection in formalin-fixed paraffin-embedded tissues of patients with lung cancer using RNA-based in situ hybridization: A preliminary feasibility study.Thorac Cancer. 2019 Oct;10(10):1936-1944. doi: 10.1111/1759-7714.13169. Epub 2019 Aug 13. Thorac Cancer. 2019. PMID: 31407509 Free PMC article.
References
-
- Mahmood R, Mason I. In-situ hybridization of radioactive riboprobes to RNA in tissue sections. Methods in Molecular Biology. 2008;461:675–686. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous