Altered patterns of global protein synthesis and translational fidelity in RPS15-mutated chronic lymphocytic leukemia
- PMID: 30181176
- PMCID: PMC6410914
- DOI: 10.1182/blood-2017-09-804401
Altered patterns of global protein synthesis and translational fidelity in RPS15-mutated chronic lymphocytic leukemia
Abstract
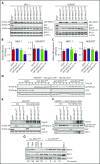
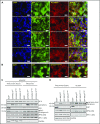
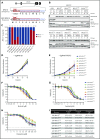
Genomic studies have recently identified RPS15 as a new driver gene in aggressive and chemorefractory cases of chronic lymphocytic leukemia (CLL). RPS15 encodes a ribosomal protein whose conserved C-terminal domain extends into the decoding center of the ribosome. We demonstrate that mutations in highly conserved residues of this domain affect protein stability, by increasing its ubiquitin-mediated degradation, and cell-proliferation rates. On the other hand, we show that mutated RPS15 can be loaded into the ribosomes, directly impacting on global protein synthesis and/or translational fidelity in a mutation-specific manner. Quantitative mass spectrometry analyses suggest that RPS15 variants may induce additional alterations in the translational machinery, as well as a metabolic shift at the proteome level in HEK293T and MEC-1 cells. These results indicate that CLL-related RPS15 mutations might act following patterns known for other ribosomal diseases, likely switching from a hypo- to a hyperproliferative phenotype driven by mutated ribosomes. In this scenario, loss of translational fidelity causing altered cell proteostasis can be proposed as a new molecular mechanism involved in CLL pathobiology.
© 2018 by The American Society of Hematology.
Conflict of interest statement
Conflict-of-interest disclosure: The authors declare no competing financial interests.
Figures



Comment in
-
Not so lost in translation: RPS15 mutations in CLL.Blood. 2018 Nov 29;132(22):2317-2319. doi: 10.1182/blood-2018-09-875179. Blood. 2018. PMID: 30498067 No abstract available.
References
-
- Fabbri G, Dalla-Favera R. The molecular pathogenesis of chronic lymphocytic leukaemia. Nat Rev Cancer. 2016;16(3):145-162. - PubMed
-
- Hamblin TJ, Davis Z, Gardiner A, Oscier DG, Stevenson FK. Unmutated Ig V(H) genes are associated with a more aggressive form of chronic lymphocytic leukemia. Blood. 1999;94(6):1848-1854. - PubMed
-
- Damle RN, Wasil T, Fais F, et al. Ig V gene mutation status and CD38 expression as novel prognostic indicators in chronic lymphocytic leukemia. Blood. 1999;94(6):1840-1847. - PubMed
-
- Sutton LA, Rosenquist R. The complex interplay between cell-intrinsic and cell-extrinsic factors driving the evolution of chronic lymphocytic leukemia. Semin Cancer Biol. 2015;34:22-35. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

