Local enrichment of HP1alpha at telomeres alters their structure and regulation of telomere protection
- PMID: 30181605
- PMCID: PMC6123478
- DOI: 10.1038/s41467-018-05840-y
Local enrichment of HP1alpha at telomeres alters their structure and regulation of telomere protection
Abstract
Enhanced telomere maintenance is evident in malignant cancers. While telomeres are thought to be inherently heterochromatic, detailed mechanisms of how epigenetic modifications impact telomere protection and structures are largely unknown in human cancers. Here we develop a molecular tethering approach to experimentally enrich heterochromatin protein HP1α specifically at telomeres. This results in increased deposition of H3K9me3 at cancer cell telomeres. Telomere extension by telomerase is attenuated, and damage-induced foci at telomeres are reduced, indicating augmentation of telomere stability. Super-resolution STORM imaging shows an unexpected increase in irregularity of telomeric structure. Telomere-tethered chromo shadow domain (CSD) mutant I165A of HP1α abrogates both the inhibition of telomere extension and the irregularity of telomeric structure, suggesting the involvement of at least one HP1α-ligand in mediating these effects. This work presents an approach to specifically manipulate the epigenetic status locally at telomeres to uncover insights into molecular mechanisms underlying telomere structural dynamics.
Conflict of interest statement
The authors declare no competing interests.
Figures
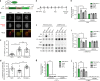



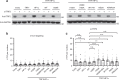


Similar articles
-
TERRA RNA binding to TRF2 facilitates heterochromatin formation and ORC recruitment at telomeres.Mol Cell. 2009 Aug 28;35(4):403-13. doi: 10.1016/j.molcel.2009.06.025. Mol Cell. 2009. PMID: 19716786 Free PMC article.
-
Inflammatory factor receptor Toll-like receptor 4 controls telomeres through heterochromatin protein 1 isoforms in liver cancer stem cell.J Cell Mol Med. 2018 Jun;22(6):3246-3258. doi: 10.1111/jcmm.13606. Epub 2018 Mar 30. J Cell Mol Med. 2018. PMID: 29602239 Free PMC article.
-
Dynamics of protein binding to telomeres in living cells: implications for telomere structure and function.Mol Cell Biol. 2004 Jun;24(12):5587-94. doi: 10.1128/MCB.24.12.5587-5594.2004. Mol Cell Biol. 2004. PMID: 15169917 Free PMC article.
-
Nucleosome organization and chromatin dynamics in telomeres.Biomol Concepts. 2015 Mar;6(1):67-75. doi: 10.1515/bmc-2014-0035. Biomol Concepts. 2015. PMID: 25720088 Review.
-
Beyond the histone tale: HP1α deregulation in breast cancer epigenetics.Cancer Biol Ther. 2015;16(2):189-200. doi: 10.1080/15384047.2014.1001277. Cancer Biol Ther. 2015. PMID: 25588111 Free PMC article. Review.
Cited by
-
In Situ Detection of Complex DNA Damage Using Microscopy: A Rough Road Ahead.Cancers (Basel). 2020 Nov 6;12(11):3288. doi: 10.3390/cancers12113288. Cancers (Basel). 2020. PMID: 33172046 Free PMC article. Review.
-
Cell Biology of Giant Cell Tumour of Bone: Crosstalk between m/wt Nucleosome H3.3, Telomeres and Osteoclastogenesis.Cancers (Basel). 2021 Oct 13;13(20):5119. doi: 10.3390/cancers13205119. Cancers (Basel). 2021. PMID: 34680268 Free PMC article. Review.
-
Human telomerase is directly regulated by non-telomeric TRF2-G-quadruplex interaction.Cell Rep. 2021 May 18;35(7):109154. doi: 10.1016/j.celrep.2021.109154. Cell Rep. 2021. PMID: 34010660 Free PMC article.
-
hTERT Increases TRF2 to Induce Telomere Compaction and Extend Cell Replicative Lifespan.Aging Cell. 2025 Aug;24(8):e70105. doi: 10.1111/acel.70105. Epub 2025 May 15. Aging Cell. 2025. PMID: 40371663 Free PMC article.
-
Regulation of Gene Expression by Telomere Position Effect.Int J Mol Sci. 2021 Nov 26;22(23):12807. doi: 10.3390/ijms222312807. Int J Mol Sci. 2021. PMID: 34884608 Free PMC article. Review.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

