Identification of key pathways and genes in response to trastuzumab treatment in breast cancer using bioinformatics analysis
- PMID: 30181805
- PMCID: PMC6114942
- DOI: 10.18632/oncotarget.24605
Identification of key pathways and genes in response to trastuzumab treatment in breast cancer using bioinformatics analysis
Abstract
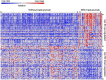
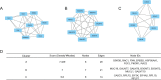
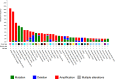
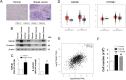
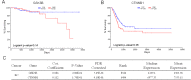
Breast cancer (BC) is one of the leading causes of death among women worldwide. The gene expression profile GSE22358 was downloaded from the Gene Expression Omnibus (GEO) database, which included 154 operable early-stage breast cancer samples treated with neoadjuvant capecitabine plus docetaxel, with (34) or without trastuzumab (120), to identify gene signatures during trastuzumab treatment and uncover their potential mechanisms. The gene ontology (GO) and Kyoto Encyclopedia of Genes and Genomes pathway (KEGG) enrichment analyses were performed, and a protein-protein interaction (PPI) network of the differentially expressed genes (DEGs) was constructed by Cytoscape software. There were 2284 DEGs, including 1231 up-regulated genes enriched in DNA replication, protein N-linked glycosylation via asparagine, and response to toxic substances, while 1053 down-regulated genes were enriched in axon guidance, protein localization to plasma membrane, protein stabilization, and protein glycosylation. Eight hub genes were identified from the PPI network, including GSK3B, RAC1, PXN, ERBB2, HSP90AA1, FGF2, PIK3R1 and RAC2. Our experimental results showed that GSK3B was also highly expressed in breast cancer tissues and was associated with poor survival, as was β-catenin. In conclusion, the present study indicated that the identified DEGs and hub genes further our understanding of the molecular mechanisms underlying trastuzumab treatment in BC and highlighted GSK3B, which might be used as a molecular target for the treatment of BC.
Keywords: bioinformatics analysis; breast cancer; differentially expressed gene; microarray; trastuzumab.
Conflict of interest statement
CONFLICTS OF INTEREST The authors declare that there are no conflicts of interest.
Figures
Similar articles
-
Identification of key pathways and genes in colorectal cancer using bioinformatics analysis.Med Oncol. 2016 Oct;33(10):111. doi: 10.1007/s12032-016-0829-6. Epub 2016 Aug 31. Med Oncol. 2016. PMID: 27581154
-
The identification of key genes and pathways in hepatocellular carcinoma by bioinformatics analysis of high-throughput data.Med Oncol. 2017 Jun;34(6):101. doi: 10.1007/s12032-017-0963-9. Epub 2017 Apr 21. Med Oncol. 2017. PMID: 28432618 Free PMC article.
-
Identification of differentially expressed genes regulated by molecular signature in breast cancer-associated fibroblasts by bioinformatics analysis.Arch Gynecol Obstet. 2018 Jan;297(1):161-183. doi: 10.1007/s00404-017-4562-y. Epub 2017 Oct 23. Arch Gynecol Obstet. 2018. PMID: 29063236
-
Identification of breast cancer hub genes and analysis of prognostic values using integrated bioinformatics analysis.Cancer Biomark. 2017 Dec 12;21(1):373-381. doi: 10.3233/CBM-170550. Cancer Biomark. 2017. PMID: 29081411
-
Assessment of the SRC Inhibition Role in the Efficacy of Breast Cancer Radiotherapy.J Lasers Med Sci. 2019 Fall;10(Suppl 1):S18-S22. doi: 10.15171/jlms.2019.S4. Epub 2019 Dec 1. J Lasers Med Sci. 2019. PMID: 32021668 Free PMC article. Review.
Cited by
-
Robust Identification of Differential Gene Expression Patterns from Multiple Transcriptomics Datasets for Early Diagnosis, Prognosis, and Therapies for Breast Cancer.Medicina (Kaunas). 2023 Sep 24;59(10):1705. doi: 10.3390/medicina59101705. Medicina (Kaunas). 2023. PMID: 37893423 Free PMC article.
-
Identification of Potential Key Genes Associated With the Pathogenesis, Metastasis, and Prognosis of Triple-Negative Breast Cancer on the Basis of Integrated Bioinformatics Analysis.Front Oncol. 2020 Jun 12;10:856. doi: 10.3389/fonc.2020.00856. eCollection 2020. Front Oncol. 2020. PMID: 32596149 Free PMC article.
-
KEGG-expressed genes and pathways in triple negative breast cancer: Protocol for a systematic review and data mining.Medicine (Baltimore). 2020 May;99(18):e19986. doi: 10.1097/MD.0000000000019986. Medicine (Baltimore). 2020. PMID: 32358373 Free PMC article.
-
Highly heterogeneous-related genes of triple-negative breast cancer: potential diagnostic and prognostic biomarkers.BMC Cancer. 2021 May 31;21(1):644. doi: 10.1186/s12885-021-08318-1. BMC Cancer. 2021. PMID: 34053447 Free PMC article.
-
Temporal dynamics from phosphoproteomics using endoscopic biopsy specimens provides new therapeutic targets in stage IV gastric cancer.Sci Rep. 2022 Mar 25;12(1):4419. doi: 10.1038/s41598-022-08430-7. Sci Rep. 2022. PMID: 35338158 Free PMC article.
References
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

