Clinical and veterinary trypanocidal benzoxaboroles target CPSF3
- PMID: 30185555
- PMCID: PMC6156652
- DOI: 10.1073/pnas.1807915115
Clinical and veterinary trypanocidal benzoxaboroles target CPSF3
Abstract
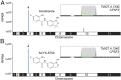
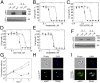
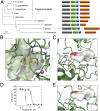
African trypanosomes cause lethal and neglected tropical diseases, known as sleeping sickness in humans and nagana in animals. Current therapies are limited, but fortunately, promising therapies are in advanced clinical and veterinary development, including acoziborole (AN5568 or SCYX-7158) and AN11736, respectively. These benzoxaboroles will likely be key to the World Health Organization's target of disease control by 2030. Their mode of action was previously unknown. We have developed a high-coverage overexpression library and use it here to explore drug mode of action in Trypanosoma brucei Initially, an inhibitor with a known target was used to select for drug resistance and to test massive parallel library screening and genome-wide mapping; this effectively identified the known target and validated the approach. Subsequently, the overexpression screening approach was used to identify the target of the benzoxaboroles, Cleavage and Polyadenylation Specificity Factor 3 (CPSF3, Tb927.4.1340). We validated the CPSF3 endonuclease as the target, using independent overexpression strains. Knockdown provided genetic validation of CPSF3 as essential, and GFP tagging confirmed the expected nuclear localization. Molecular docking and CRISPR-Cas9-based editing demonstrated how acoziborole can specifically block the active site and mRNA processing by parasite, but not host CPSF3. Thus, our findings provide both genetic and chemical validation for CPSF3 as an important drug target in trypanosomes and reveal inhibition of mRNA maturation as the mode of action of the trypanocidal benzoxaboroles. Understanding the mechanism of action of benzoxaborole-based therapies can assist development of improved therapies, as well as the prediction and monitoring of resistance, if or when it arises.
Keywords: CPSF73; N-myristoyltransferase; Ysh1; drug discovery; genetic screening.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

References
-
- Fairlamb AH, Horn D. Melarsoprol resistance in African trypanosomiasis. Trends Parasitol. 2018;34:481–492. - PubMed
-
- Simarro PP, Franco J, Diarra A, Postigo JA, Jannin J. Update on field use of the available drugs for the chemotherapy of human African trypanosomiasis. Parasitology. 2012;139:842–846. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

