Protein-protein interactions in "cis-AT" polyketide synthases
- PMID: 30188553
- PMCID: PMC6207950
- DOI: 10.1039/c8np00058a
Protein-protein interactions in "cis-AT" polyketide synthases
Abstract
Covering: up to the end of 2018 Polyketides are a valuable source of bioactive and clinically important molecules. The biosynthesis of these chemically complex molecules has led to the discovery of equally complex polyketide synthase (PKS) pathways. Crystallography has yielded snapshots of individual catalytic domains, di-domains, and multi-domains from a variety of PKS megasynthases, and cryo-EM studies have provided initial views of a PKS module in a series of defined biochemical states. Here, we review the structural and biochemical results that shed light on the protein-protein interactions critical to catalysis by PKS systems with an embedded acyltransferase. Interactions include those that occur both within and between PKS modules, as well as with accessory enzymes.
Figures
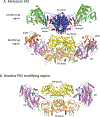





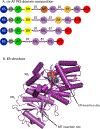
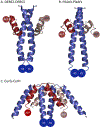
References
-
- Newman DJ and Cragg GM, J. Nat. Prod, 2016, 79, 629–661. - PubMed
-
- Cane DE, Walsh CT and Khosla C, Science, 1998, 282, 63–68. - PubMed
-
- Jenke-Kodama H, Sandmann A, Muller R and Dittmann E, Mol. Biol. Evol, 2005, 22, 2027–2039. - PubMed
-
- Staunton J and Weissman KJ, Nat. Prod. Rep, 2001, 18, 380–416. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

