Genome-wide identification and characterization of LRR-RLKs reveal functional conservation of the SIF subfamily in cotton (Gossypium hirsutum)
- PMID: 30189845
- PMCID: PMC6128003
- DOI: 10.1186/s12870-018-1395-1
Genome-wide identification and characterization of LRR-RLKs reveal functional conservation of the SIF subfamily in cotton (Gossypium hirsutum)
Abstract
Background: As one of the largest subfamilies of the receptor-like protein kinases (RLKs) in plants, Leucine Rich Repeats-RLKs (LRR-RLKs) are involved in many critical biological processes including growth, development and stress responses in addition to various physiological roles. Arabidopsis contains 234 LRR-RLKs, and four members of Stress Induced Factor (SIF) subfamily (AtSIF1-AtSIF4) which are involved in abiotic and biotic stress responses. Herein, we aimed at identification and functional characterization of SIF subfamily in cultivated tetraploid cotton Gossypium hirsutum.
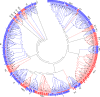
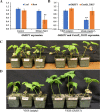
Results: Genome-wide analysis of cotton LRR-RLK gene family identified 543 members and phylogenetic analysis led to the identification of 6 cotton LRR-RLKs with high homology to Arabidopsis SIFs. Of the six SIF homologs, GhSIF1 is highly conserved exhibiting 46-47% of homology with AtSIF subfamily in amino acid sequence. The GhSIF1 was transiently silenced using Virus-Induced Gene Silencing system specifically targeting the 3' Untranslated Region. The transiently silenced cotton seedlings showed enhanced salt tolerance compared to the control plants. Further, the transiently silenced plants showed better growth, lower electrolyte leakage, and higher chlorophyll and biomass contents.
Conclusions: Overall, 543 LRR-RLK genes were identified using genome-wide analysis in cultivated tetraploid cotton G. hirsutum. The present investigation also demonstrated the conserved salt tolerance function of SIF family member in cotton. The GhSIF1 gene can be knocked out using genome editing technologies to improve salt tolerance in cotton.
Keywords: Genome-wide analysis; Gossypium hirsutum; LRR-RLKs; Salt tolerance.
Conflict of interest statement
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Figures



References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

