Differences in antigenic sites and other functional regions between genotype A and G mumps virus surface proteins
- PMID: 30190529
- PMCID: PMC6127219
- DOI: 10.1038/s41598-018-31630-z
Differences in antigenic sites and other functional regions between genotype A and G mumps virus surface proteins
Erratum in
-
Publisher Correction: Differences in antigenic sites and other functional regions between genotype A and G mumps virus surface proteins.Sci Rep. 2020 Mar 20;10(1):5456. doi: 10.1038/s41598-020-62301-7. Sci Rep. 2020. PMID: 32198462 Free PMC article.
Abstract
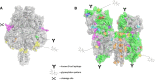
The surface proteins of the mumps virus, the fusion protein (F) and haemagglutinin-neuraminidase (HN), are key factors in mumps pathogenesis and are important targets for the immune response during mumps virus infection. We compared the predicted amino acid sequences of the F and HN genes from Dutch mumps virus samples from the pre-vaccine era (1957-1982) with mumps virus genotype G strains (from 2004 onwards). Genotype G is the most frequently detected mumps genotype in recent outbreaks in vaccinated communities, especially in Western Europe, the USA and Japan. Amino acid differences between the Jeryl Lynn vaccine strains (genotype A) and genotype G strains were predominantly located in known B-cell epitopes and in N-linked glycosylation sites on the HN protein. There were eight variable amino acid positions specific to genotype A or genotype G sequences in five known B-cell epitopes of the HN protein. These differences may account for the reported antigenic differences between Jeryl Lynn and genotype G strains. We also found amino acid differences in and near sites on the HN protein that have been reported to play a role in mumps virus pathogenesis. These differences may contribute to the occurrence of genotype G outbreaks in vaccinated communities.
Conflict of interest statement
The authors declare no competing interests.
Figures



Similar articles
-
Variability of hemagglutinin-neuraminidase and nucleocapsid protein of vaccine and wild-type mumps virus strains.Infect Genet Evol. 2008 Sep;8(5):603-13. doi: 10.1016/j.meegid.2008.04.007. Epub 2008 Apr 22. Infect Genet Evol. 2008. PMID: 18508415
-
Serologic Cross-Reactivity between the Mumps Virus Vaccine Genotype A Strain and the Circulating Genotype G Strain.Viruses. 2024 Sep 8;16(9):1434. doi: 10.3390/v16091434. Viruses. 2024. PMID: 39339910 Free PMC article.
-
Structure-based design of glycoprotein subunit vaccines for mumps.Proc Natl Acad Sci U S A. 2024 Nov 19;121(47):e2404053121. doi: 10.1073/pnas.2404053121. Epub 2024 Nov 11. Proc Natl Acad Sci U S A. 2024. PMID: 39527740 Free PMC article.
-
Exploring the Mumps Virus Glycoproteins: A Review.Viruses. 2022 Jun 18;14(6):1335. doi: 10.3390/v14061335. Viruses. 2022. PMID: 35746805 Free PMC article. Review.
-
Genomic diversity of mumps virus and global distribution of the 12 genotypes.Rev Med Virol. 2015 Mar;25(2):85-101. doi: 10.1002/rmv.1819. Epub 2014 Nov 26. Rev Med Virol. 2015. PMID: 25424978 Review.
Cited by
-
Combining genomics and epidemiology to track mumps virus transmission in the United States.PLoS Biol. 2020 Feb 11;18(2):e3000611. doi: 10.1371/journal.pbio.3000611. eCollection 2020 Feb. PLoS Biol. 2020. PMID: 32045407 Free PMC article.
-
Genomic characterization of mumps viruses from a large-scale mumps outbreak in Arkansas, 2016.Infect Genet Evol. 2019 Nov;75:103965. doi: 10.1016/j.meegid.2019.103965. Epub 2019 Jul 15. Infect Genet Evol. 2019. PMID: 31319177 Free PMC article.
-
Detecting critical slowing down in high-dimensional epidemiological systems.PLoS Comput Biol. 2020 Mar 9;16(3):e1007679. doi: 10.1371/journal.pcbi.1007679. eCollection 2020 Mar. PLoS Comput Biol. 2020. PMID: 32150536 Free PMC article.
-
Immunoinformatics approach for predicting epitopes in HN and F proteins of Porcine rubulavirus.PLoS One. 2020 Sep 25;15(9):e0239785. doi: 10.1371/journal.pone.0239785. eCollection 2020. PLoS One. 2020. PMID: 32976525 Free PMC article.
-
Investigating Local Patterns of Mumps Virus Circulation, Using a Combination of Molecular Tools.Viruses. 2023 Dec 13;15(12):2420. doi: 10.3390/v15122420. Viruses. 2023. PMID: 38140661 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

