Integrative detection and analysis of structural variation in cancer genomes
- PMID: 30202056
- PMCID: PMC6301019
- DOI: 10.1038/s41588-018-0195-8
Integrative detection and analysis of structural variation in cancer genomes
Abstract
Structural variants (SVs) can contribute to oncogenesis through a variety of mechanisms. Despite their importance, the identification of SVs in cancer genomes remains challenging. Here, we present a framework that integrates optical mapping, high-throughput chromosome conformation capture (Hi-C), and whole-genome sequencing to systematically detect SVs in a variety of normal or cancer samples and cell lines. We identify the unique strengths of each method and demonstrate that only integrative approaches can comprehensively identify SVs in the genome. By combining Hi-C and optical mapping, we resolve complex SVs and phase multiple SV events to a single haplotype. Furthermore, we observe widespread structural variation events affecting the functions of noncoding sequences, including the deletion of distal regulatory sequences, alteration of DNA replication timing, and the creation of novel three-dimensional chromatin structural domains. Our results indicate that noncoding SVs may be underappreciated mutational drivers in cancer genomes.
Figures

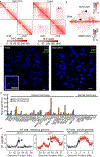



References
-
- Hanahan D & Weinberg RA Hallmarks of cancer: the next generation. Cell 144, 646–674 (2011). - PubMed
-
- Soda M et al. Identification of the transforming EML4-ALK fusion gene in non-small-cell lung cancer. Nature 448, 561–566 (2007). - PubMed
-
- Rowley JD Letter: A new consistent chromosomal abnormality in chronic myelogenous leukaemia identified by quinacrine fluorescence and Giemsa staining. Nature 243, 290–293 (1973). - PubMed
Publication types
MeSH terms
Substances
Grants and funding
- 615584/ERC_/European Research Council/International
- R01 HG009906/HG/NHGRI NIH HHS/United States
- R01 HG003143/HG/NHGRI NIH HHS/United States
- U54 DK107980/DK/NIDDK NIH HHS/United States
- P01 GM085354/GM/NIGMS NIH HHS/United States
- U01 CA200060/CA/NCI NIH HHS/United States
- U54 DK107965/DK/NIDDK NIH HHS/United States
- DP5 OD023071/OD/NIH HHS/United States
- 202878/Z/16/Z/WT_/Wellcome Trust/United Kingdom
- 20412/CRUK_/Cancer Research UK/United Kingdom
- R24 DK106766/DK/NIDDK NIH HHS/United States
- 22398/CRUK_/Cancer Research UK/United Kingdom
- R01 GM083337/GM/NIGMS NIH HHS/United States
- U54 HG004592/HG/NHGRI NIH HHS/United States
- R35 GM124820/GM/NIGMS NIH HHS/United States
- U41 HG007000/HG/NHGRI NIH HHS/United States
LinkOut - more resources
Full Text Sources
Other Literature Sources

