Zebrafish expression reporters and mutants reveal that the IgSF cell adhesion molecule Dscamb is required for feeding and survival
- PMID: 30204029
- PMCID: PMC6464818
- DOI: 10.1080/01677063.2018.1493479
Zebrafish expression reporters and mutants reveal that the IgSF cell adhesion molecule Dscamb is required for feeding and survival
Abstract
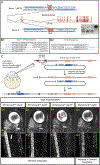
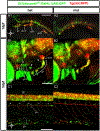
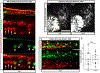
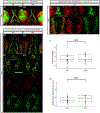
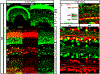
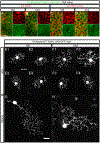
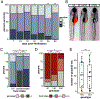
Down syndrome cell adhesion molecules (DSCAMs) are broadly expressed in nervous systems and play conserved roles in programmed cell death, neuronal migration, axon guidance, neurite branching and spacing, and synaptic targeting. However, DSCAMs appear to have distinct functions in different vertebrate animals, and little is known about their functions outside the retina. We leveraged the genetic tractability and optical accessibility of larval zebrafish to investigate the expression and function of a DSCAM family member, dscamb. Using targeted genome editing to create transgenic reporters and loss-of-function mutant alleles, we discovered that dscamb is expressed broadly throughout the brain, spinal cord, and peripheral nervous system, but is not required for overall structural organization of the brain. Despite the absence of obvious anatomical defects, homozygous dscamb mutants were deficient in their ability to ingest food and rarely survived to adulthood. Thus, we have discovered a novel function for dscamb in feeding behavior. The mutant and transgenic lines generated in these studies will provide valuable tools for identifying the molecular and cellular bases of these behaviors.
Keywords: Dscam; Zebrafish; behavior; expression pattern; feeding; genome editing.
Figures

Similar articles
-
Dscamb regulates cone mosaic formation in zebrafish via filopodium-mediated homotypic recognition.Nat Commun. 2025 Mar 25;16(1):2501. doi: 10.1038/s41467-025-57506-1. Nat Commun. 2025. PMID: 40133281 Free PMC article.
-
Expression patterns of dscam and sdk gene paralogs in developing zebrafish retina.Mol Vis. 2018 Jul 19;24:443-458. eCollection 2018. Mol Vis. 2018. PMID: 30078982 Free PMC article.
-
Localization of Cadm2a and Cadm3 proteins during development of the zebrafish nervous system.J Comp Neurol. 2011 Aug 1;519(11):2252-70. doi: 10.1002/cne.22627. J Comp Neurol. 2011. PMID: 21456004
-
Roles for DSCAM and DSCAML1 in central nervous system development and disease.Adv Neurobiol. 2014;8:249-70. doi: 10.1007/978-1-4614-8090-7_11. Adv Neurobiol. 2014. PMID: 25300140 Review.
-
Molecular diversity of Dscam and self-recognition.Adv Exp Med Biol. 2012;739:262-75. doi: 10.1007/978-1-4614-1704-0_17. Adv Exp Med Biol. 2012. PMID: 22399408 Review.
Cited by
-
Keratins and the plakin family cytolinker proteins control the length of epithelial microridge protrusions.Elife. 2020 Sep 7;9:e58149. doi: 10.7554/eLife.58149. Elife. 2020. PMID: 32894222 Free PMC article.
-
From Biomphalaria glabrata to Drosophila melanogaster and Anopheles gambiae: the diversity and role of FREPs and Dscams in immune response.Front Immunol. 2025 Apr 30;16:1579905. doi: 10.3389/fimmu.2025.1579905. eCollection 2025. Front Immunol. 2025. PMID: 40370466 Free PMC article. Review.
-
Zebrafish Feed Intake: A Systematic Review for Standardizing Feeding Management in Laboratory Conditions.Biology (Basel). 2024 Mar 23;13(4):209. doi: 10.3390/biology13040209. Biology (Basel). 2024. PMID: 38666821 Free PMC article. Review.
-
Structure and dynamics of liposomes designed for drug delivery: coarse-grained molecular dynamics simulations to reveal the role of lipopolymer incorporation.RSC Adv. 2020 Jan 22;10(7):3745-3755. doi: 10.1039/c9ra08632c. eCollection 2020 Jan 22. RSC Adv. 2020. PMID: 35492626 Free PMC article.
-
Dscamb regulates cone mosaic formation in zebrafish via filopodium-mediated homotypic recognition.Nat Commun. 2025 Mar 25;16(1):2501. doi: 10.1038/s41467-025-57506-1. Nat Commun. 2025. PMID: 40133281 Free PMC article.
References
-
- Agarwala KL, Ganesh S, Amano K, Suzuki T, & Yamakawa K (2001). DSCAM, a Highly Conserved Gene in Mammals, Expressed in Differentiating Mouse Brain. Biochemical and Biophysical Research Communications, 281(3), 697–705. - PubMed
-
- Agarwala KL, Ganesh S, Tsutsumi Y, Suzuki T, Amano K, & Yamakawa K (2001). Cloning and functional characterization of DSCAML1, a novel DSCAM-like cell adhesion molecule that mediates homophilic intercellular adhesion. Biochemical and Biophysical Research Communications, 285(3), 760–772. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
