Transmissible ER stress reconfigures the AML bone marrow compartment
- PMID: 30206307
- PMCID: PMC6411460
- DOI: 10.1038/s41375-018-0254-2
Transmissible ER stress reconfigures the AML bone marrow compartment
Abstract
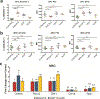
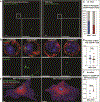
Successive adaptation of the bone marrow (BM) from homeostatic hematopoietic microenvironment to a self-reinforcing niche is an integral aspect of leukemogenesis. Yet, the cellular mechanisms underlying these functional alterations remain to be defined. Here, we found that AML incursion precipitates compartmental endoplasmic reticulum (ER) stress and an unfolded protein response (UPR) in both leukemia and stromal cells. We observed that extracellular vesicles (EV) transmit ER stress in vivo from the AML xenograft to BM stroma, whereby the upregulation of core UPR components drives subsequent osteolineage differentiation of mesenchymal stem cells (MSC). Finally, we show that the underlying mechanism involves quantitative incorporation and cell-cell transfer of Bone Morphogenic Protein 2 (BMP2), a potent osteogenic signal, by AML-EVs. Corroborative studies in AML patient samples support the translational relevance of AML-EVs as a platform for BMP trafficking and source of compartmental crosstalk. Transmissible ER stress was previously identified as a source of chemoresistance in solid tumor models, and this work reveals a role in remodeling the BM niche in AML.
Conflict of interest statement
Figures




Similar articles
-
Acute myeloid leukemia transforms the bone marrow niche into a leukemia-permissive microenvironment through exosome secretion.Leukemia. 2018 Mar;32(3):575-587. doi: 10.1038/leu.2017.259. Epub 2017 Aug 17. Leukemia. 2018. PMID: 28816238 Free PMC article.
-
Mesenchymal niche remodeling impairs hematopoiesis via stanniocalcin 1 in acute myeloid leukemia.J Clin Invest. 2020 Jun 1;130(6):3038-3050. doi: 10.1172/JCI133187. J Clin Invest. 2020. PMID: 32364536 Free PMC article.
-
Abnormal adipogenic signaling in the bone marrow mesenchymal stem cells contributes to supportive microenvironment for leukemia development.Cell Commun Signal. 2023 Oct 10;21(1):277. doi: 10.1186/s12964-023-01231-z. Cell Commun Signal. 2023. PMID: 37817179 Free PMC article.
-
Transforming the Niche: The Emerging Role of Extracellular Vesicles in Acute Myeloid Leukaemia Progression.Int J Mol Sci. 2024 Apr 17;25(8):4430. doi: 10.3390/ijms25084430. Int J Mol Sci. 2024. PMID: 38674015 Free PMC article. Review.
-
Extracellular vesicles secreted by leukemic cells as mediators of dysregulated hematopoiesis: acute myeloid leukemia as a case in point.Expert Rev Hematol. 2025 Jan-Mar;18(3):225-237. doi: 10.1080/17474086.2025.2471860. Epub 2025 Feb 26. Expert Rev Hematol. 2025. PMID: 40008450 Review.
Cited by
-
Salubrinal Exposes Anticancer Properties in Inflammatory Breast Cancer Cells by Manipulating the Endoplasmic Reticulum Stress Pathway.Front Oncol. 2021 May 20;11:654940. doi: 10.3389/fonc.2021.654940. eCollection 2021. Front Oncol. 2021. PMID: 34094947 Free PMC article.
-
Interplay between endoplasmic reticulum stress and non-coding RNAs in cancer.J Hematol Oncol. 2020 Dec 2;13(1):163. doi: 10.1186/s13045-020-01002-0. J Hematol Oncol. 2020. PMID: 33267910 Free PMC article. Review.
-
Bone marrow niches in myelodysplastic syndromes.J Cancer Metastasis Treat. 2021;7:52. doi: 10.20517/2394-4722.2021.120. Epub 2021 Jul 14. J Cancer Metastasis Treat. 2021. PMID: 34746416 Free PMC article.
-
Deciphering Tumor Niches: Lessons From Solid and Hematological Malignancies.Front Immunol. 2021 Nov 10;12:766275. doi: 10.3389/fimmu.2021.766275. eCollection 2021. Front Immunol. 2021. PMID: 34858421 Free PMC article. Review.
-
Unfolded Protein Response in Leukemia: From Basic Understanding to Therapeutic Opportunities.Trends Cancer. 2020 Nov;6(11):960-973. doi: 10.1016/j.trecan.2020.05.012. Epub 2020 Jun 13. Trends Cancer. 2020. PMID: 32540455 Free PMC article. Review.
References
-
- El-Badri NS, Wang BY, Cherry Good RA. Osteoblasts promote engraftment of allogeneic hematopoietic stem cells. Exp Hematol. 1998;26(2):110–6. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

