Developmental processes regulate craniofacial variation in disease and evolution
- PMID: 30207415
- PMCID: PMC6349473
- DOI: 10.1002/dvg.23249
Developmental processes regulate craniofacial variation in disease and evolution
Abstract
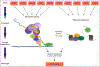
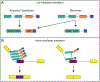
Variation in development mediates phenotypic differences observed in evolution and disease. Although the mechanisms underlying phenotypic variation are still largely unknown, recent research suggests that variation in developmental processes may play a key role. Developmental processes mediate genotype-phenotype relationships and consequently play an important role regulating phenotypes. In this review, we provide an example of how shared and interacting developmental processes may explain convergence of phenotypes in spliceosomopathies and ribosomopathies. These data also suggest a shared pathway to disease treatment. We then discuss three major mechanisms that contribute to variation in developmental processes: genetic background (gene-gene interactions), gene-environment interactions, and developmental stochasticity. Finally, we comment on evolutionary alterations to developmental processes, and the evolution of disease buffering mechanisms.
Keywords: craniofacial anomalies; evolution of development; genotype-phenotype relationships; morphological variation; ribosomopathies; spliceosomopathies.
© 2018 Wiley Periodicals, Inc.
Figures

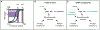
Similar articles
-
Developmental mechanisms underlying variation in craniofacial disease and evolution.Dev Biol. 2016 Jul 15;415(2):188-197. doi: 10.1016/j.ydbio.2015.12.019. Epub 2015 Dec 24. Dev Biol. 2016. PMID: 26724698 Review.
-
From genotype to phenotype: the differential expression of FGF, FGFR, and TGFbeta genes characterizes human cranioskeletal development and reflects clinical presentation in FGFR syndromes.Plast Reconstr Surg. 2001 Dec;108(7):2026-39; discussion 2040-6. doi: 10.1097/00006534-200112000-00030. Plast Reconstr Surg. 2001. PMID: 11743396
-
Developmental interactions and the constituents of quantitative variation.Evolution. 2001 Feb;55(2):232-45. doi: 10.1111/j.0014-3820.2001.tb01289.x. Evolution. 2001. PMID: 11308082
-
Craniofacial variation and developmental divergence in primate and human evolution.Novartis Found Symp. 2007;284:262-73; discussion 273-9. doi: 10.1002/9780470319390.ch17. Novartis Found Symp. 2007. PMID: 17710858 Review.
-
Isolating and quantifying the role of developmental noise in generating phenotypic variation.PLoS Comput Biol. 2019 Apr 22;15(4):e1006943. doi: 10.1371/journal.pcbi.1006943. eCollection 2019 Apr. PLoS Comput Biol. 2019. PMID: 31009449 Free PMC article.
Cited by
-
Pharmacological Inhibition of the Spliceosome SF3b Complex by Pladienolide-B Elicits Craniofacial Developmental Defects in Mouse and Zebrafish.Birth Defects Res. 2024 Nov;116(11):e2404. doi: 10.1002/bdr2.2404. Birth Defects Res. 2024. PMID: 39494782 Free PMC article.
-
Regional variability in craniofacial stiffness: a study in normal and Crouzon mice during postnatal development.Biomech Model Mechanobiol. 2025 Aug;24(4):1207-1222. doi: 10.1007/s10237-025-01962-7. Epub 2025 May 25. Biomech Model Mechanobiol. 2025. PMID: 40413698 Free PMC article.
-
Satb2 regulates proliferation and nuclear integrity of pre-osteoblasts.Bone. 2019 Oct;127:488-498. doi: 10.1016/j.bone.2019.07.017. Epub 2019 Jul 17. Bone. 2019. PMID: 31325654 Free PMC article.
-
Variation in phenotypes from a Bmp-Gata3 genetic pathway is modulated by Shh signaling.PLoS Genet. 2021 May 25;17(5):e1009579. doi: 10.1371/journal.pgen.1009579. eCollection 2021 May. PLoS Genet. 2021. PMID: 34033651 Free PMC article.
-
Integrated Transcriptome and Network Analysis Reveals Spatiotemporal Dynamics of Calvarial Suturogenesis.Cell Rep. 2020 Jul 7;32(1):107871. doi: 10.1016/j.celrep.2020.107871. Cell Rep. 2020. PMID: 32640236 Free PMC article.
References
-
- Allende-Vega N, Dayal S, Agarwala U, Sparks A, Bourdon JC, Saville MK. 2013. p53 is activated in response to disruption of the pre-mRNA splicing machinery. Oncogene 32: 1–14. - PubMed
-
- Ameyar-Zazoua M, Rachez C, Souidi M, Robin P, Fritsch L, Young R, Morozova N, Fenouil R, Descostes N, Andrau JC, Mathieu J, Hamiche A, Ait-Si-Ali S, Muchardt C, Batsche E, Harel-Bellan A. 2012. Argonaute proteins couple chromatin silencing to alternative splicing. Nat Struct Mol Biol 19: 998–1004. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

