MIEF1 Microprotein Regulates Mitochondrial Translation
- PMID: 30215512
- PMCID: PMC6443411
- DOI: 10.1021/acs.biochem.8b00726
MIEF1 Microprotein Regulates Mitochondrial Translation
Abstract
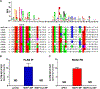
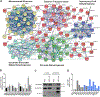
Recent technological advances led to the discovery of hundreds to thousands of peptides and small proteins (microproteins) encoded by small open reading frames (smORFs). Characterization of new microproteins demonstrates their role in fundamental biological processes and highlights the value in discovering and characterizing more microproteins. The elucidation of microprotein-protein interactions (MPIs) is useful for determining the biochemical and cellular roles of microproteins. In this study, we characterize the protein interaction partners of mitochondrial elongation factor 1 microprotein (MIEF1-MP) using a proximity labeling strategy that relies on APEX2. MIEF1-MP localizes to the mitochondrial matrix where it interacts with the mitochondrial ribosome (mitoribosome). Functional studies demonstrate that MIEF1-MP regulates mitochondrial translation via its binding to the mitoribosome. Loss of MIEF1-MP decreases the mitochondrial translation rate, while an elevated level of MIEF1-MP increases the translation rate. The identification of MIEF1-MP reveals a new gene involved in this process.
Figures


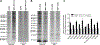
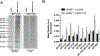
Similar articles
-
An Inner Mitochondrial Membrane Microprotein from the SLC35A4 Upstream ORF Regulates Cellular Metabolism.J Mol Biol. 2024 May 15;436(10):168559. doi: 10.1016/j.jmb.2024.168559. Epub 2024 Apr 3. J Mol Biol. 2024. PMID: 38580077 Free PMC article.
-
Human MIEF1 recruits Drp1 to mitochondrial outer membranes and promotes mitochondrial fusion rather than fission.EMBO J. 2011 Jun 24;30(14):2762-78. doi: 10.1038/emboj.2011.198. EMBO J. 2011. PMID: 21701560 Free PMC article.
-
The mitochondrial elongation factors MIEF1 and MIEF2 exert partially distinct functions in mitochondrial dynamics.Exp Cell Res. 2013 Nov 1;319(18):2893-904. doi: 10.1016/j.yexcr.2013.07.010. Epub 2013 Jul 20. Exp Cell Res. 2013. PMID: 23880462
-
Mitochondrial-derived microproteins: from discovery to function.Trends Genet. 2025 Feb;41(2):132-145. doi: 10.1016/j.tig.2024.11.010. Epub 2024 Dec 16. Trends Genet. 2025. PMID: 39690001 Free PMC article. Review.
-
Peptides derived from small mitochondrial open reading frames: Genomic, biological, and therapeutic implications.Exp Cell Res. 2020 Aug 15;393(2):112056. doi: 10.1016/j.yexcr.2020.112056. Epub 2020 May 6. Exp Cell Res. 2020. PMID: 32387288 Free PMC article. Review.
Cited by
-
Small open reading frames and cellular stress responses.Mol Omics. 2019 Apr 1;15(2):108-116. doi: 10.1039/c8mo00283e. Epub 2019 Feb 27. Mol Omics. 2019. PMID: 30810554 Free PMC article. Review.
-
The Role of Mitochondria-Derived Peptides in Cardiovascular Diseases and Their Potential as Therapeutic Targets.Int J Mol Sci. 2021 Aug 16;22(16):8770. doi: 10.3390/ijms22168770. Int J Mol Sci. 2021. PMID: 34445477 Free PMC article. Review.
-
Noncanonical altPIDD1 protein: unveiling the true major translational output of the PIDD1 gene.Life Sci Alliance. 2024 Nov 12;8(2):e202402910. doi: 10.26508/lsa.202402910. Print 2025 Feb. Life Sci Alliance. 2024. PMID: 39532532 Free PMC article.
-
Casting CRISPR-Cas13d to fish for microprotein functions in animal development.iScience. 2022 Nov 10;25(12):105547. doi: 10.1016/j.isci.2022.105547. eCollection 2022 Dec 22. iScience. 2022. PMID: 36444300 Free PMC article. Review.
-
Accurate annotation of human protein-coding small open reading frames.Nat Chem Biol. 2020 Apr;16(4):458-468. doi: 10.1038/s41589-019-0425-0. Epub 2019 Dec 9. Nat Chem Biol. 2020. PMID: 31819274 Free PMC article.
References
-
- Burge CB, and Karlin S (1998) Finding the genes in genomic DNA, Current opinion in structural biology 8, 346–354. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

