In vivo cross-linking supports a head-to-tail mechanism for regulation of the plant plasma membrane P-type H+-ATPase
- PMID: 30217814
- PMCID: PMC6222091
- DOI: 10.1074/jbc.RA118.003528
In vivo cross-linking supports a head-to-tail mechanism for regulation of the plant plasma membrane P-type H+-ATPase
Abstract
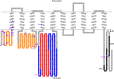
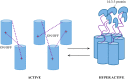
In higher plants, a P-type proton-pumping ATPase generates the proton-motive force essential for the function of all other transporters and for proper growth and development. X-ray crystallographic studies of the plant plasma membrane proton pump have provided information on amino acids involved in ATP catalysis but provided no information on the structure of the C-terminal regulatory domain. Despite progress in elucidating enzymes involved in the signaling pathways that activate or inhibit this pump, the site of interaction of the C-terminal regulatory domain with the catalytic domains remains a mystery. Genetic studies have pointed to amino acids in various parts of the protein that may be involved, but direct chemical evidence for which ones are specifically interacting with the C terminus is lacking. In this study, we used in vivo cross-linking experiments with a photoreactive unnatural amino acid, p-benzoylphenylalanine, and tandem MS to obtain direct evidence that the C-terminal regulatory domain interacts with amino acids located within the N-terminal actuator domain. Our observations are consistent with a mechanism in which intermolecular, rather than intramolecular, interactions are involved. Our model invokes a "head-to-tail" organization of ATPase monomers in which the C-terminal domain of one ATPase molecule interacts with the actuator domain of another ATPase molecule. This model serves to explain why cross-linked peptides are found only in dimers and trimers, and it is consistent with prior studies suggesting that within the membrane the protein can be organized as homopolymers, including dimers, trimers, and hexamers.
Keywords: Arabidopsis thaliana; H+-ATPase; mass spectrometry (MS); protein cross-linking; protein-protein interaction.
© 2018 Nguyen et al.
Conflict of interest statement
The authors declare that they have no conflicts of interest with the contents of this article
Figures



References
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

