Large-scale investigation of the reasons why potentially important genes are ignored
- PMID: 30226837
- PMCID: PMC6143198
- DOI: 10.1371/journal.pbio.2006643
Large-scale investigation of the reasons why potentially important genes are ignored
Abstract
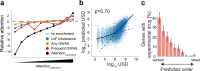
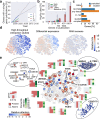
Biomedical research has been previously reported to primarily focus on a minority of all known genes. Here, we demonstrate that these differences in attention can be explained, to a large extent, exclusively from a small set of identifiable chemical, physical, and biological properties of genes. Together with knowledge about homologous genes from model organisms, these features allow us to accurately predict the number of publications on individual human genes, the year of their first report, the levels of funding awarded by the National Institutes of Health (NIH), and the development of drugs against disease-associated genes. By explicitly identifying the reasons for gene-specific bias and performing a meta-analysis of existing computational and experimental knowledge bases, we describe gene-specific strategies for the identification of important but hitherto ignored genes that can open novel directions for future investigation.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures


Comment in
-
Reply to "Far away from the lamppost".PLoS Biol. 2018 Dec 11;16(12):e3000075. doi: 10.1371/journal.pbio.3000075. eCollection 2018 Dec. PLoS Biol. 2018. PMID: 30532190 Free PMC article.
-
Far away from the lamppost.PLoS Biol. 2018 Dec 11;16(12):e3000067. doi: 10.1371/journal.pbio.3000067. eCollection 2018 Dec. PLoS Biol. 2018. PMID: 30532236 Free PMC article.
References
-
- Hoffmann R, Valencia A. Life cycles of successful genes. Trends Genet. 2003;19(2):79–81. Epub 2003/01/28. . - PubMed
-
- Gans Joshua MF, Stern Scott. Patents, Papers, Pairs & Secrets: Contracting over the disclosure of scientific knowledge. Statement is only present in self-hosted early draft: http://fmurray.scripts.mit.edu/docs/Gans.Murray.Stern%20_KnowledgeDisclo.... 2008 [cited 2018 Aug 22].
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases

