Analysis of bZIP Transcription Factor Family and Their Expressions under Salt Stress in Chlamydomonas reinhardtii
- PMID: 30227676
- PMCID: PMC6164503
- DOI: 10.3390/ijms19092800
Analysis of bZIP Transcription Factor Family and Their Expressions under Salt Stress in Chlamydomonas reinhardtii
Abstract
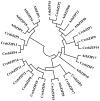
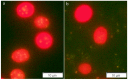
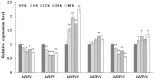
The basic leucine-region zipper (bZIP) transcription factors (TFs) act as crucial regulators in various biological processes and stress responses in plants. Currently, bZIP family members and their functions remain elusive in the green unicellular algae Chlamydomonas reinhardtii, an important model organism for molecular investigation with genetic engineering aimed at increasing lipid yields for better biodiesel production. In this study, a total of 17 C. reinhardtii bZIP (CrebZIP) TFs containing typical bZIP structure were identified by a genome-wide analysis. Analysis of the CrebZIP protein physicochemical properties, phylogenetic tree, conserved domain, and secondary structure were conducted. CrebZIP gene structures and their chromosomal assignment were also analyzed. Physiological and photosynthetic characteristics of C. reinhardtii under salt stress were exhibited as lower cell growth and weaker photosynthesis, but increased lipid accumulation. Meanwhile, the expression profiles of six CrebZIP genes were induced to change significantly during salt stress, indicating that certain CrebZIPs may play important roles in mediating photosynthesis and lipid accumulation of microalgae in response to stresses. The present work provided a valuable foundation for functional dissection of CrebZIPs, benefiting the development of better strategies to engineer the regulatory network in microalgae for enhancing biofuel and biomass production.
Keywords: Chlamydomonas reinhardtii; bZIP transcription factors; lipid accumulation; photosynthesis; salt stress; transcriptional regulation.
Conflict of interest statement
The authors declare no conflict of interest.
Figures






References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

