Genetic and scRNA-seq Analysis Reveals Distinct Cell Populations that Contribute to Salivary Gland Development and Maintenance
- PMID: 30232460
- PMCID: PMC6145895
- DOI: 10.1038/s41598-018-32343-z
Genetic and scRNA-seq Analysis Reveals Distinct Cell Populations that Contribute to Salivary Gland Development and Maintenance
Abstract
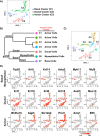
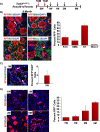
Stem and progenitor cells of the submandibular salivary gland (SMG) give rise to, maintain, and regenerate the multiple lineages of mature epithelial cells including those belonging to the ductal, acinar, basal and myoepithelial subtypes. Here we have exploited single cell RNA-sequencing and in vivo genetic lineage tracing technologies to generate a detailed map of the cell fate trajectories and branch points of the basal and myoepithelial cell populations of the mouse SMG during embryonic development and in adults. Our studies show that the transcription factor p63 and alpha-smooth muscle actin (SMA) serve as faithful markers of the basal and myoepithelial cell lineages, respectively and that both cell types are endowed with progenitor cell properties. However, p63+ basal and SMA+ myoepithelial cells exhibit distinct cell fates by virtue of maintaining different cellular lineages during morphogenesis and in adults. Collectively, our results reveal the dynamic and complex nature of the diverse SMG cell populations and highlight the distinct differentiation potential of the p63 and SMA expressing subtypes in the stem and progenitor cell hierarchy. Long term these findings have profound implications towards a better understanding of the molecular mechanisms that dictate lineage commitment and differentiation programs during development and adult gland maintenance.
Conflict of interest statement
The authors declare no competing interests.
Figures






References
Publication types
MeSH terms
Substances
Grants and funding
- R25 GM095459/GM/NIGMS NIH HHS/United States
- T32 DE023526/DE/NIDCR NIH HHS/United States
- R01 DE027660/DE/NIDCR NIH HHS/United States
- DE023526/U.S. Department of Health & Human Services | NIH | National Institute of Dental and Craniofacial Research (NIDCR)/International
- DE025889/U.S. Department of Health & Human Services | NIH | National Institute of Dental and Craniofacial Research (NIDCR)/International
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

