Quaternary structure of the small amino acid transporter OprG from Pseudomonas aeruginosa
- PMID: 30237175
- PMCID: PMC6222097
- DOI: 10.1074/jbc.RA118.004461
Quaternary structure of the small amino acid transporter OprG from Pseudomonas aeruginosa
Abstract
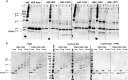
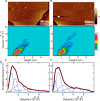
Pseudomonas aeruginosa is an opportunistic human pathogen that causes nosocomial infections. The P. aeruginosa outer membrane contains specific porins that enable substrate uptake, with the outer membrane protein OprG facilitating transport of small, uncharged amino acids. However, the pore size of an eight-stranded β-barrel monomer of OprG is too narrow to accommodate even the smallest transported amino acid, glycine, raising the question of how OprG facilitates amino acid uptake. Pro-92 of OprG is critically important for amino acid transport, with a P92A substitution inhibiting transport and the NMR structure of this variant revealing that this substitution produces structural changes in the barrel rim and restricts loop motions. OprG may assemble into oligomers in the outer membrane (OM) whose subunit interfaces could form a transport channel. Here, we explored the contributions of the oligomeric state and the extracellular loops to OprG's function. Using chemical cross-linking to determine the oligomeric structures of both WT and P92A OprG in native outer membranes and atomic force microscopy, and single-molecule fluorescence of the purified proteins reconstituted into lipid bilayers, we found that both protein variants form oligomers, supporting the notion that subunit interfaces in the oligomer could provide a pathway for amino acid transport. Furthermore, performing transport assays with loop-deleted OprG variants, we found that these variants also can transport small amino acids, indicating that the loops are not solely responsible for substrate transport. We propose that OprG functions as an oligomer and that conformational changes in the barrel-loop region might be crucial for its activity.
Keywords: Pseudomonas aeruginosa (P. aeruginosa); atomic force microscopy (AFM); chemical modification; fluorescence; membrane protein; outer membrane; single-molecule biophysics; transporter.
© 2018 Sanganna Gari et al.
Conflict of interest statement
The authors declare that they have no conflicts of interest with the contents of this article
Figures



References
-
- Kang C. I., Kim S. H., Kim H. B., Park S. W., Choe Y. J., Oh M. D., Kim E. C., and Choe K. W. (2003) Pseudomonas aeruginosa bacteremia: risk factors for mortality and influence of delayed receipt of effective antimicrobial therapy on clinical outcome. Clin. Infect. Dis. 37, 745–751 10.1086/377200 - DOI - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

