PLK4 is a microtubule-associated protein that self-assembles promoting de novo MTOC formation
- PMID: 30237222
- PMCID: PMC6398482
- DOI: 10.1242/jcs.219501
PLK4 is a microtubule-associated protein that self-assembles promoting de novo MTOC formation
Abstract
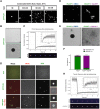
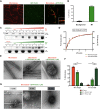
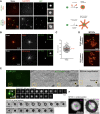
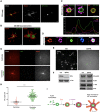
The centrosome is an important microtubule-organising centre (MTOC) in animal cells. It consists of two barrel-shaped structures, the centrioles, surrounded by the pericentriolar material (PCM), which nucleates microtubules. Centrosomes can form close to an existing structure (canonical duplication) or de novo How centrosomes form de novo is not known. The master driver of centrosome biogenesis, PLK4, is critical for the recruitment of several centriole components. Here, we investigate the beginning of centrosome biogenesis, taking advantage of Xenopus egg extracts, where PLK4 can induce de novo MTOC formation ( Eckerdt et al., 2011; Zitouni et al., 2016). Surprisingly, we observe that in vitro, PLK4 can self-assemble into condensates that recruit α- and β-tubulins. In Xenopus extracts, PLK4 assemblies additionally recruit STIL, a substrate of PLK4, and the microtubule nucleator γ-tubulin, forming acentriolar MTOCs de novo The assembly of these robust microtubule asters is independent of dynein, similar to what is found for centrosomes. We suggest a new mechanism of action for PLK4, where it forms a self-organising catalytic scaffold that recruits centriole components, PCM factors and α- and β-tubulins, leading to MTOC formation.This article has an associated First Person interview with the first author of the paper.
Keywords: Centrosome; De novo assembly; In vitro reconstitution; MTOCs; Microtubule nucleation; PCM; PLK4; Supramolecular assembly.
© 2018. Published by The Company of Biologists Ltd.
Conflict of interest statement
Competing interestsThe authors declare no competing or financial interests.
Figures
Similar articles
-
Asterless is a scaffold for the onset of centriole assembly.Nature. 2010 Oct 7;467(7316):714-8. doi: 10.1038/nature09445. Epub 2010 Sep 19. Nature. 2010. PMID: 20852615
-
Self-assembly of pericentriolar material in interphase cells lacking centrioles.Elife. 2022 Jul 5;11:e77892. doi: 10.7554/eLife.77892. Elife. 2022. PMID: 35787744 Free PMC article.
-
Plk4 triggers autonomous de novo centriole biogenesis and maturation.J Cell Biol. 2021 May 3;220(5):e202008090. doi: 10.1083/jcb.202008090. J Cell Biol. 2021. PMID: 33760919 Free PMC article.
-
Polar expeditions--provisioning the centrosome for mitosis.Nat Cell Biol. 2003 Jun;5(6):505-11. doi: 10.1038/ncb0603-505. Nat Cell Biol. 2003. PMID: 12776127 Review.
-
Bridging centrioles and PCM in proper space and time.Essays Biochem. 2018 Dec 7;62(6):793-801. doi: 10.1042/EBC20180036. Print 2018 Dec 7. Essays Biochem. 2018. PMID: 30429283 Free PMC article. Review.
Cited by
-
A self-assembled cylindrical platform for Plk4-induced centriole biogenesis.Open Biol. 2020 Aug;10(8):200102. doi: 10.1098/rsob.200102. Epub 2020 Aug 19. Open Biol. 2020. PMID: 32810424 Free PMC article. Review.
-
Multidomain ribosomal protein trees and the planctobacterial origin of neomura (eukaryotes, archaebacteria).Protoplasma. 2020 May;257(3):621-753. doi: 10.1007/s00709-019-01442-7. Epub 2020 Jan 3. Protoplasma. 2020. PMID: 31900730 Free PMC article. Review.
-
PLK4 self-phosphorylation drives the selection of a single site for procentriole assembly.J Cell Biol. 2023 Dec 4;222(12):e202301069. doi: 10.1083/jcb.202301069. Epub 2023 Sep 29. J Cell Biol. 2023. PMID: 37773039 Free PMC article.
-
TRIM37 prevents formation of centriolar protein assemblies by regulating Centrobin.Elife. 2021 Jan 25;10:e62640. doi: 10.7554/eLife.62640. Elife. 2021. PMID: 33491649 Free PMC article.
-
Daughter centrioles assemble preferentially towards the nuclear envelope in Drosophila syncytial embryos.Open Biol. 2022 Jan;12(1):210343. doi: 10.1098/rsob.210343. Epub 2022 Jan 19. Open Biol. 2022. PMID: 35042404 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

