Notch2 controls non-autonomous Wnt-signalling in chronic lymphocytic leukaemia
- PMID: 30242258
- PMCID: PMC6155045
- DOI: 10.1038/s41467-018-06069-5
Notch2 controls non-autonomous Wnt-signalling in chronic lymphocytic leukaemia
Abstract
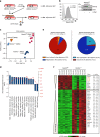
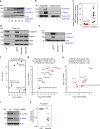
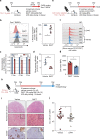
The Wnt signalling pathway, one of the core de-regulated pathways in chronic lymphocytic leukaemia (CLL), is activated in only a subset of patients through somatic mutations. Here we describe alternative, microenvironment-dependent mechanisms of Wnt activation in malignant B cells. We show that tumour cells specifically induce Notch2 activity in mesenchymal stromal cells (MSCs) required for the transcription of the complement factor C1q. MSC-derived C1q in turn inhibits Gsk3-β mediated degradation of β-catenin in CLL cells. Additionally, stromal Notch2 activity regulates N-cadherin expression in CLL cells, which interacts with and further stabilises β-catenin. Together, these stroma Notch2-dependent mechanisms induce strong activation of canonical Wnt signalling in CLL cells. Pharmacological inhibition of the Wnt pathway impairs microenvironment-mediated survival of tumour cells. Similarly, inhibition of Notch signalling diminishes survival of stroma-protected CLL cells in vitro and disease engraftment in vivo. Notch2 activation in the microenvironment is a pre-requisite for the activation of canonical Wnt signalling in tumour cells.
Conflict of interest statement
The authors declare no competing interests.
Figures





References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

