Binge Ethanol Drinking Produces Sexually Divergent and Distinct Changes in Nucleus Accumbens Signaling Cascades and Pathways in Adult C57BL/6J Mice
- PMID: 30250478
- PMCID: PMC6139464
- DOI: 10.3389/fgene.2018.00325
Binge Ethanol Drinking Produces Sexually Divergent and Distinct Changes in Nucleus Accumbens Signaling Cascades and Pathways in Adult C57BL/6J Mice
Abstract
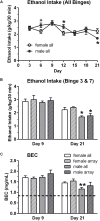
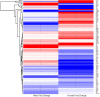
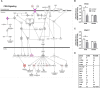
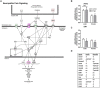
We previously determined that repeated binge ethanol drinking produced sex differences in the regulation of signaling downstream of Group 1 metabotropic glutamate receptors in the nucleus accumbens (NAc) of adult C57BL/6J mice. The purpose of the present study was to characterize RNA expression differences in the NAc of adult male and female C57BL/6J mice following 7 binge ethanol drinking sessions, when compared with controls consuming water. This binge drinking procedure produced high intakes (average >2.2 g/kg/30 min) and blood ethanol concentrations (average >1.3 mg/ml). Mice were euthanized at 24 h after the 7th binge session, and focused qPCR array analysis was employed on NAc tissue to quantify expression levels of 384 genes in a customized Mouse Mood Disorder array, with a focus on glutamatergic signaling (3 arrays/group). We identified significant regulation of 50 genes in male mice and 70 genes in female mice after 7 ethanol binges. Notably, 14 genes were regulated in both males and females, representing common targets to binge ethanol drinking. However, expression of 10 of these 14 genes was strongly dimorphic (e.g., opposite regulation for genes such as Crhr2, Fos, Nos1, and Star), and only 4 of the 14 genes were regulated in the same direction (Drd5, Grm4, Ranbp9, and Reln). Interestingly, the top 30 regulated genes by binge ethanol drinking for each sex differed markedly in the male and female mice, and this divergent neuroadaptive response in the NAc could result in dysregulation of distinct biological pathways between the sexes. Characterization of the expression differences with Ingenuity Pathway Analysis was used to identify Canonical Pathways, Upstream Regulators, and significant Biological Functions. Expression differences suggested that hormone signaling and immune function were altered by binge drinking in female mice, whereas neurotransmitter metabolism was a central target of binge ethanol drinking in male mice. Thus, these results indicate that the transcriptional response to repeated binge ethanol drinking was strongly influenced by sex, and they emphasize the importance of considering sex in the development of potential pharmacotherapeutic targets for the treatment of alcohol use disorder.
Keywords: C57BL/6J mice; alcohol; hormone signaling; immune function; neurotransmitter metabolism; qPCR arrays; sex differences.
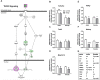
Figures
Similar articles
-
Functional regulation of PI3K-associated signaling in the accumbens by binge alcohol drinking in male but not female mice.Neuropharmacology. 2016 Jun;105:164-174. doi: 10.1016/j.neuropharm.2016.01.010. Epub 2016 Jan 7. Neuropharmacology. 2016. PMID: 26773198 Free PMC article.
-
Effects of chemogenetic manipulation of the nucleus accumbens core in male C57BL/6J mice.Alcohol. 2021 Mar;91:21-27. doi: 10.1016/j.alcohol.2020.10.005. Epub 2020 Nov 4. Alcohol. 2021. PMID: 33160072 Free PMC article.
-
Sex differences in the synergistic effect of prior binge drinking and traumatic stress on subsequent ethanol intake and neurochemical responses in adult C57BL/6J mice.Alcohol. 2018 Sep;71:33-45. doi: 10.1016/j.alcohol.2018.02.004. Epub 2018 Mar 3. Alcohol. 2018. PMID: 29966824 Free PMC article.
-
Protein Kinase C Epsilon Activity in the Nucleus Accumbens and Central Nucleus of the Amygdala Mediates Binge Alcohol Consumption.Biol Psychiatry. 2016 Mar 15;79(6):443-51. doi: 10.1016/j.biopsych.2015.01.019. Epub 2015 Mar 5. Biol Psychiatry. 2016. PMID: 25861702 Free PMC article.
-
Stress and gonadal steroid influences on alcohol drinking and withdrawal, with focus on animal models in females.Front Neuroendocrinol. 2023 Oct;71:101094. doi: 10.1016/j.yfrne.2023.101094. Epub 2023 Aug 8. Front Neuroendocrinol. 2023. PMID: 37558184 Free PMC article. Review.
Cited by
-
Sex differences in nucleus accumbens core circuitry engaged by binge-like ethanol drinking.bioRxiv [Preprint]. 2024 Aug 19:2024.08.15.608144. doi: 10.1101/2024.08.15.608144. bioRxiv. 2024. Update in: Psychopharmacology (Berl). 2025 Aug 22. doi: 10.1007/s00213-025-06875-y. PMID: 39229134 Free PMC article. Updated. Preprint.
-
Impact of Genetic Variation on Stress-Related Ethanol Consumption.Alcohol Clin Exp Res. 2019 Jul;43(7):1391-1402. doi: 10.1111/acer.14073. Epub 2019 May 21. Alcohol Clin Exp Res. 2019. PMID: 31034606 Free PMC article.
-
Blood and brain gene expression signatures of chronic intermittent ethanol consumption in mice.PLoS Comput Biol. 2022 Feb 17;18(2):e1009800. doi: 10.1371/journal.pcbi.1009800. eCollection 2022 Feb. PLoS Comput Biol. 2022. PMID: 35176017 Free PMC article.
-
Housing Condition Differentially Impacts Escalation of Alcohol Intake, Relapse-Like Drinking, Anxiety-Like Behavior, and Stress History Effects by Sex.Alcohol Clin Exp Res. 2021 Feb;45(2):480-489. doi: 10.1111/acer.14540. Epub 2021 Jan 20. Alcohol Clin Exp Res. 2021. PMID: 33351976 Free PMC article.
-
Cocaine Regulates NLRP3 Inflammasome Activity and CRF Signaling in a Region- and Sex-Dependent Manner in Rat Brain.Biomedicines. 2023 Jun 23;11(7):1800. doi: 10.3390/biomedicines11071800. Biomedicines. 2023. PMID: 37509440 Free PMC article.
References
-
- Bell R. L., Hauser S. R., McClintick J., Rahman S., Edenberg H. J., Szumlinski K. K., et al. (2016). Ethanol-associated changes in glutamate reward neurocircuitry: A minireview of clinical and preclinical genetic findings. Prog. Mol. Biol. Transl. Sci. 137 41–85. 10.1016/bs.pmbts.2015.10.018 - DOI - PMC - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

