Population structure of a vector of human diseases: Aedes aegypti in its ancestral range, Africa
- PMID: 30250667
- PMCID: PMC6145026
- DOI: 10.1002/ece3.4278
Population structure of a vector of human diseases: Aedes aegypti in its ancestral range, Africa
Abstract
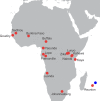
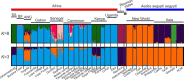
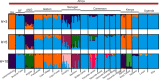
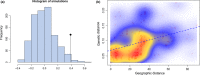
Aedes aegypti, the major vector of dengue, yellow fever, chikungunya, and Zika viruses, remains of great medical and public health concern. There is little doubt that the ancestral home of the species is Africa. This mosquito invaded the New World 400-500 years ago and later, Asia. However, little is known about the genetic structure and history of Ae. aegypti across Africa, as well as the possible origin(s) of the New World invasion. Here, we use ~17,000 genome-wide single nucleotide polymorphisms (SNPs) to characterize a heretofore undocumented complex picture of this mosquito across its ancestral range in Africa. We find signatures of human-assisted migrations, connectivity across long distances in sylvan populations, and of local admixture between domestic and sylvan populations. Finally, through a phylogenetic analysis combined with the genetic structure analyses, we suggest West Africa and especially Angola as the source of the New World's invasion, a scenario that fits well with the historic record of 16th-century slave trade between Africa and Americas.
Keywords: Aedes aegypti; Africa; SNP‐chip; genetics; migration; population structure.
Figures



References
-
- Bennett, K. L. , Shija, F. , Linton, Y. M. , Misinzo, G. , Kaddumukasa, M. , Djouaka, R. , … Walton, C. (2016). Historical environmental change in Africa drives divergence and admixture of Aedes aegypti mosquitoes: A precursor to successful worldwide colonization? Molecular Ecology, 25, 4337–4354. 10.1111/mec.13762 - DOI - PubMed
-
- Bosio, C. F. , Harrington, L. C. , Jones, J. W. , Sithiprasana, R. , Norris, D. E. , & Scott, T. W. (2005). Genetic structure of Aedes aegypti populations in Thailand using mitochondrial DNA. American Journal of Tropical Medicine and Hygiene, 72, 434–442. - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

