Comprehensive manipulation of glycosylation profiles across development scales
- PMID: 30252592
- PMCID: PMC6380396
- DOI: 10.1080/19420862.2018.1527665
Comprehensive manipulation of glycosylation profiles across development scales
Abstract
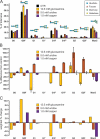
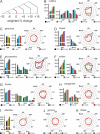
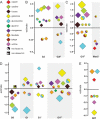
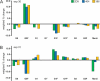
The extent and pattern of glycosylation on therapeutic antibodies can influence their circulatory half-life, engagement of effector functions, and immunogenicity, with direct consequences to efficacy and patient safety. Hence, controlling glycosylation patterns is central to any drug development program, yet poses a formidable challenge to the bio-manufacturing industry. Process changes, which can affect glycosylation patterns, range from manufacturing at different scales or sites, to switching production process mode, all the way to using alternative host cell lines. In the emerging space of biosimilars development, often times all of these aspects apply. Gaining a deep understanding of the direction and extent to which glycosylation quality attributes can be modulated is key for efficient fine-tuning of glycan profiles in a stage appropriate manner, but establishment of such platform knowledge is time consuming and resource intensive. Here we report an inexpensive and highly adaptable screening system for comprehensive modulation of glycans on antibodies expressed in CHO cells. We characterize 10 media additives in univariable studies and in combination, using a design of experiments approach to map the design space for tuning glycosylation profile attributes. We introduce a robust workflow that does not require automation, yet enables rapid process optimization. We demonstrate scalability across deep wells, shake flasks, AMBR-15 cell culture system, and 2 L single-use bioreactors. Further, we show that it is broadly applicable to different molecules and host cell lineages. This universal approach permits fine-tuned modulation of glycan product quality, reduces development costs, and enables agile implementation of process changes throughout the product lifecycle.
Keywords: ADC; DoE; feed additives; glycosylation; media development; multivariate analysis; product quality; scalability.
Figures


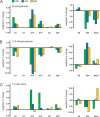
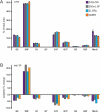
References
-
- Shields RL, Lai J, Keck R, O’Connell LY, Hong K, Gloria Meng Y, Weikert SHA, Presta LG. Lack of fucose on human IgG1 N-linked oligosaccharide improves binding to human FcγRIII and antibody-dependent cellular toxicity. J Biol Chem. 2002;277(30):26733–26740. doi: 10.1074/jbc.M202069200. - DOI - PubMed
-
- Shinkawa T, Nakamura K, Yamane N, Shoji-Hosaka E, Kanda Y, Sakurada M, Uchida K, Anazawa H, Satoh M, Yamasaki M, et al. The absence of fucose but not the presence of galactose or bisecting N-acetylglucosamine of human IgG1 complex-type oligosaccharides shows the critical role of enhancing antibody-dependent cellular cytotoxicity. J Biol Chem. 2003;278(5):3466–3473. doi: 10.1074/jbc.M210665200. - DOI - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
