Chlorate Specifically Targets Oxidant-Starved, Antibiotic-Tolerant Populations of Pseudomonas aeruginosa Biofilms
- PMID: 30254119
- PMCID: PMC6156191
- DOI: 10.1128/mBio.01400-18
Chlorate Specifically Targets Oxidant-Starved, Antibiotic-Tolerant Populations of Pseudomonas aeruginosa Biofilms
Abstract
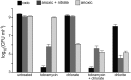
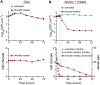
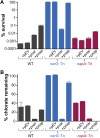
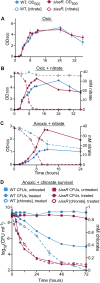
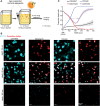
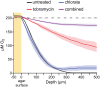
Nitrate respiration is a widespread mode of anaerobic energy generation used by many bacterial pathogens, and the respiratory nitrate reductase, Nar, has long been known to reduce chlorate to the toxic oxidizing agent chlorite. Here, we demonstrate the antibacterial activity of chlorate against Pseudomonas aeruginosa, a representative pathogen that can inhabit hypoxic or anoxic host microenvironments during infection. Aerobically grown P. aeruginosa cells are tobramycin sensitive but chlorate tolerant. In the absence of oxygen or an alternative electron acceptor, cells are tobramycin tolerant but chlorate sensitive via Nar-dependent reduction. The fact that chlorite, the product of chlorate reduction, is not detected in culture supernatants suggests that it may react rapidly and be retained intracellularly. Tobramycin and chlorate target distinct populations within metabolically stratified aggregate biofilms; tobramycin kills cells on the oxic periphery, whereas chlorate kills hypoxic and anoxic cells in the interior. In a matrix populated by multiple aggregates, tobramycin-mediated death of surface aggregates enables deeper oxygen penetration into the matrix, benefiting select aggregate populations by increasing survival and removing chlorate sensitivity. Finally, lasR mutants, which commonly arise in P. aeruginosa infections and are known to withstand conventional antibiotic treatment, are hypersensitive to chlorate. A lasR mutant shows a propensity to respire nitrate and reduce chlorate more rapidly than the wild type does, consistent with its heightened chlorate sensitivity. These findings illustrate chlorate's potential to selectively target oxidant-starved pathogens, including physiological states and genotypes of P. aeruginosa that represent antibiotic-tolerant populations during infections.IMPORTANCE The anaerobic growth and survival of bacteria are often correlated with physiological tolerance to conventional antibiotics, motivating the development of novel strategies targeting pathogens in anoxic environments. A key challenge is to identify drug targets that are specific to this metabolic state. Chlorate is a nontoxic compound that can be reduced to toxic chlorite by a widespread enzyme of anaerobic metabolism. We tested the antibacterial properties of chlorate against Pseudomonas aeruginosa, a pathogen that can inhabit hypoxic or anoxic microenvironments, including those that arise in human infection. Chlorate and the antibiotic tobramycin kill distinct metabolic populations in P. aeruginosa biofilms, where chlorate targets anaerobic cells that tolerate tobramycin. Chlorate is particularly effective against P. aeruginosalasR mutants, which are frequently isolated from human infections and more resistant to some antibiotics. This work suggests that chlorate may hold potential as an anaerobic prodrug.
Keywords: Nar; Pseudomonas aeruginosa; antibiotic tolerance; biofilms; chlorate; nitrate reduction; prodrug.
Copyright © 2018 Spero and Newman.
Figures
Similar articles
-
Targeting Anaerobic Respiration in Pseudomonas aeruginosa with Chlorate Improves Healing of Chronic Wounds.Adv Wound Care (New Rochelle). 2024 Feb;13(2):53-69. doi: 10.1089/wound.2023.0036. Epub 2023 Aug 14. Adv Wound Care (New Rochelle). 2024. PMID: 37432895 Free PMC article.
-
Mechanisms of chlorate toxicity and resistance in Pseudomonas aeruginosa.Mol Microbiol. 2022 Oct;118(4):321-335. doi: 10.1111/mmi.14972. Epub 2022 Aug 15. Mol Microbiol. 2022. PMID: 36271736 Free PMC article.
-
Effect of oxygen limitation on the in vitro antimicrobial susceptibility of clinical isolates of Pseudomonas aeruginosa grown planktonically and as biofilms.Eur J Clin Microbiol Infect Dis. 2005 Oct;24(10):677-87. doi: 10.1007/s10096-005-0031-9. Eur J Clin Microbiol Infect Dis. 2005. PMID: 16249934
-
Pseudomonas aeruginosa hypoxic or anaerobic biofilm infections within cystic fibrosis airways.Trends Microbiol. 2009 Mar;17(3):130-8. doi: 10.1016/j.tim.2008.12.003. Epub 2009 Feb 21. Trends Microbiol. 2009. PMID: 19231190 Review.
-
Microenvironmental characteristics and physiology of biofilms in chronic infections of CF patients are strongly affected by the host immune response.APMIS. 2017 Apr;125(4):276-288. doi: 10.1111/apm.12668. APMIS. 2017. PMID: 28407427 Review.
Cited by
-
Aggregation of Nontuberculous Mycobacteria Is Regulated by Carbon-Nitrogen Balance.mBio. 2019 Aug 13;10(4):e01715-19. doi: 10.1128/mBio.01715-19. mBio. 2019. PMID: 31409683 Free PMC article.
-
Targeting Anaerobic Respiration in Pseudomonas aeruginosa with Chlorate Improves Healing of Chronic Wounds.Adv Wound Care (New Rochelle). 2024 Feb;13(2):53-69. doi: 10.1089/wound.2023.0036. Epub 2023 Aug 14. Adv Wound Care (New Rochelle). 2024. PMID: 37432895 Free PMC article.
-
Visualization of mRNA Expression in Pseudomonas aeruginosa Aggregates Reveals Spatial Patterns of Fermentative and Denitrifying Metabolism.Appl Environ Microbiol. 2022 Jun 14;88(11):e0043922. doi: 10.1128/aem.00439-22. Epub 2022 May 19. Appl Environ Microbiol. 2022. PMID: 35586988 Free PMC article.
-
From the soil to the clinic: the impact of microbial secondary metabolites on antibiotic tolerance and resistance.Nat Rev Microbiol. 2022 Mar;20(3):129-142. doi: 10.1038/s41579-021-00620-w. Epub 2021 Sep 16. Nat Rev Microbiol. 2022. PMID: 34531577 Free PMC article. Review.
-
Fungal biofilm architecture produces hypoxic microenvironments that drive antifungal resistance.Proc Natl Acad Sci U S A. 2020 Sep 8;117(36):22473-22483. doi: 10.1073/pnas.2003700117. Epub 2020 Aug 26. Proc Natl Acad Sci U S A. 2020. PMID: 32848055 Free PMC article.
References
-
- Cook GM, Greening C, Hards K, Berney M. 2014. Energetics of pathogenic bacteria and opportunities for drug development. Advances in microbial physiology. Adv Bact Pathog Biol 65:1–62. - PubMed
-
- Hassett DJ, Cuppoletti J, Trapnell B, Lymar SV, Rowe JJ, Yoon SS, Hilliard GM, Parvatiyar K, Kamani MC, Wozniak DJ, Hwang SH, McDermott TR, Ochsner UA. 2002. Anaerobic metabolism and quorum sensing by Pseudomonas aeruginosa biofilms in chronically infected cystic fibrosis airways: rethinking antibiotic treatment strategies and drug targets. Adv Drug Deliv Rev 54:1425–1443. doi:10.1016/S0169-409X(02)00152-7. - DOI - PubMed
-
- Winter SE, Winter MG, Xavier MN, Thiennimitr P, Poon V, Keestra AM, Laughlin RC, Gomez G, Wu J, Lawhon SD, Popova IE, Parikh SJ, Adams LG, Tsolis RM, Stewart VJ, Baumler AJ. 2013. Host-derived nitrate boosts growth of E. coli in the inflamed gut. Science 339:708–711. doi:10.1126/science.1232467. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

